
Concept explainers
DNA footprint protection (described in Research Technique
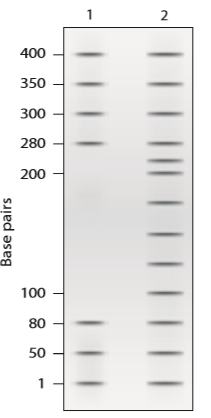
a. Explain why this gel provides evidence that the cloned DNA may act as a promoter sequence.
b. Approximately what length is the DNA region protected by RNA pol II and TFIIs?
c. What additional genetic experiments would you suggest to verify that this region of cloned DNA contains a functional promoter?
Want to see the full answer?
Check out a sample textbook solution
Chapter 8 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- Question #5: Assume that two genes are identified that confer gametophytic facultative apomixis in soybean. The genes show independent assortment. Recessive alleles at both loci are required for the facultative apomixis. Facultative apomixis is triggered when the temperature at pollination is above 20 degrees C. At temperatures below 20 degrees C, all reproduction is sexual, independent of genotype. A facultative apomict male, capable of producing viable pollen, was crossed with a sexually reproducing female. Assuming the parents are completely inbred, what are the predicted phenotypic ratios (apomict: non-apomict) for the F1, F2, and DH (F1-derived) generations at each of the following temperatures*: a) 15°C? b) 25°C? *for full credit, show crosses and genotypes where appropriate. Remember to position the female first (left side) in the cross. Type your answer here:arrow_forwarda. What percentage of a drug is eliminated after 4 half-lives? Please round to the nearest percent. b. What will happen to elimination of the drug in the previous question if the system is saturated? explain and show any math involvedarrow_forwardIf you wanted to reduce the difference between peak and trough levels that occur with repeated administration of a drug, how would you adjust the dose and dose interval without changing the plateau concentration (plateau is the average of peak and trough levels)? Select your answers for both dose and interval. Hint: It may be helpful to think about this problem using an example such as food. How would you eat if you wanted to maintain very steady hunger/satiety levels without changing your total caloric intake? Options: A. Dose; Increase dose B. Dose; Decrease dose C. Dose; Do not change dose D. Interval; Increase the interval between doses (give the drug less frequently) E. Interval; Decrease the interval between doses (give the drug more frequently) F. Interval; Do not change the intervalarrow_forward
- What percentage of a drug is eliminated after 4 half-lives? Please round to the nearest percent. Show the matharrow_forwardBriefly explain the 6 domain of interprofessional collaboration: Role clarification, Team functioning, Interprofessional communication, Patient/client/family/community-centered care, Interprofessional conflict resolution, Collaborative leadership. Provide a specific negative events that nursing student would observe in a clinical setting for each domain.arrow_forwardwhat is an intermittent water course and what kind of fish habitat it would providearrow_forward
- why are native freshwater mussels are an important part of great lakes ecosystemarrow_forwardwhat morphological features differentiate the lamprey species and other species in the great lakesarrow_forwardThere are a wide range of therapeutic applications available as options for patients. Medical professionals should be aware of these applications so they can make informed recommendations to patients. To gain a better understanding of some therapeutic applications and how they are related to RNA and mRNA, research long non-coding RNA. Respond to the following in a minimum of 175 words: What is lncRNA and what does it do? How does IncRNA differ from mRNA? What are some therapeutic applications associated with lncRNA? Think about possible future uses of this application. What are the advantages and disadvantages of this application and its continued use?arrow_forward
- four fish or mussel species that are native to the great lakesarrow_forwardThere are a wide range of therapeutic applications available as options for patients. Medical professionals should be aware of these applications so they can make informed recommendations to patients. To gain a better understanding of some therapeutic applications and how they are related to RNA and mRNA, research long non-coding RNA. Respond to the following in a minimum of 175 words: What is lncRNA and what does it do? How does IncRNA differ from mRNA? What are some therapeutic applications associated with lncRNA? Think about possible future uses of this application. What are the advantages and disadvantages of this application and its continued use?arrow_forwardfour physial characteristics of a fish or a mussel that would help you identify it to a speciesarrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning
Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage LearningCase Studies In Health Information ManagementBiologyISBN:9781337676908Author:SCHNERINGPublisher:Cengage
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage LearningCase Studies In Health Information ManagementBiologyISBN:9781337676908Author:SCHNERINGPublisher:Cengage Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning





