
To analyze:
Four introns labeled A to D are present in eukaryotic gene (
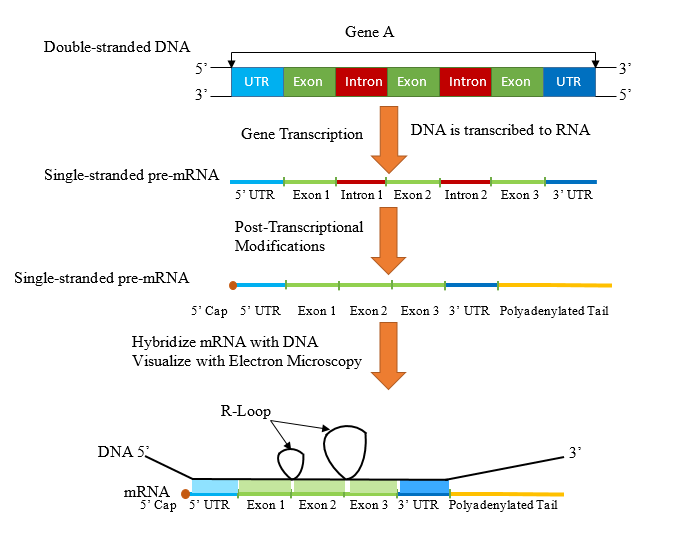
a. The R-loop structure, seen with electron microscopy is to be illustrated.
b. The introns are to be labelled.
c. Single stranded or double stranded, intron region is to be determined.
Introduction:
DNA contains coding and non-coding regions called exons and introns respectively. During post-transcriptional modifications of eukaryotic DNA, the introns are spliced out and mature mRNA is synthesized containing only exons. R-looping was first discovered and described in
Want to see the full answer?
Check out a sample textbook solution
Chapter 8 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- Amino Acid Coclow TABle 3' Gly Phe Leu (G) (F) (L) 3- Val (V) Arg (R) Ser (S) Ala (A) Lys (K) CAG G Glu Asp (E) (D) Ser (S) CCCAGUCAGUCAGUCAG 0204 C U A G C Asn (N) G 4 A AGU C GU (5) AC C UGA A G5 C CUGACUGACUGACUGAC Thr (T) Met (M) lle £€ (1) U 4 G Tyr Σε (Y) U Cys (C) C A G Trp (W) 3' U C A Leu בוט His Pro (P) ££ (H) Gin (Q) Arg 흐름 (R) (L) Start Stop 8. Transcription and Translation Practice: (Video 10-1 and 10-2) A. Below is the sense strand of a DNA gene. Using the sense strand, create the antisense DNA strand and label the 5' and 3' ends. B. Use the antisense strand that you create in part A as a template to create the mRNA transcript of the gene and label the 5' and 3' ends. C. Translate the mRNA you produced in part B into the polypeptide sequence making sure to follow all the rules of translation. 5'-AGCATGACTAATAGTTGTTGAGCTGTC-3' (sense strand) 4arrow_forwardWhat is the structure and function of Eukaryotic cells, including their organelles? How are Eukaryotic cells different than Prokaryotic cells, in terms of evolution which form of the cell might have came first? How do Eukaryotic cells become malignant (cancerous)?arrow_forwardWhat are the roles of DNA and proteins inside of the cell? What are the building blocks or molecular components of the DNA and proteins? How are proteins produced within the cell? What connection is there between DNA, proteins, and the cell cycle? What is the relationship between DNA, proteins, and Cancer?arrow_forward
- please fill in the empty sports, thank you!arrow_forwardIn one paragraph show how atoms and they're structure are related to the structure of dna and proteins. Talk about what atoms are. what they're made of, why chemical bonding is important to DNA?arrow_forwardWhat are the structure and properties of atoms and chemical bonds (especially how they relate to DNA and proteins).arrow_forward
- The Sentinel Cell: Nature’s Answer to Cancer?arrow_forwardMolecular Biology Question You are working to characterize a novel protein in mice. Analysis shows that high levels of the primary transcript that codes for this protein are found in tissue from the brain, muscle, liver, and pancreas. However, an antibody that recognizes the C-terminal portion of the protein indicates that the protein is present in brain, muscle, and liver, but not in the pancreas. What is the most likely explanation for this result?arrow_forwardMolecular Biology Explain/discuss how “slow stop” and “quick/fast stop” mutants wereused to identify different protein involved in DNA replication in E. coli.arrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning
Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning
 BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning





