
Biochemistry: Concepts and Connections (2nd Edition)
2nd Edition
ISBN: 9780134641621
Author: Dean R. Appling, Spencer J. Anthony-Cahill, Christopher K. Mathews
Publisher: PEARSON
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 8, Problem 11P
The following data describe the catalysis of cleavage of peptide bonds small peptides by the enzyme elastase.
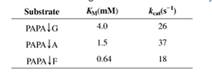
The arrow indicates the peptide bond cleaved each case.
a. If a mixture of these three substrates was presented to elastase with the concentration of each peptide equal to 0.5 mM, which would be digested most rapidly? Which most slowly? (Assume enzyme present in excess.)
b. On basis of these data, suggest what features of amino acid sequence dictate the specificity of proteolytic cleavage by elastase.
c. Elastase is closely related to chymotrypsin. Suggest two kinds of amino acid residues you might expect to find in or near the active site.
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
The reaction A+B → C + D AG°' = -7.3 kcal/mol
can be coupled with which of the following unfavorable reactions to
drive it forward?
A. EFG+HAG° = 5.6 kcal/mol.
B. J+KZ+A AG° = 2.3 kcal/mol.
C. P+RY+DAG° = 8.2 kcal/mol.
D. C + T → V + W AG°' = -5.9 kcal/mol.
E. AN→ Q+KAG°' = 4.3 kcal/mol.
What would be the toxicological endpoints for neurotoxicity?
What are "endpoints" in toxicology exactly? Please give an intuitive easy explanation
Chapter 8 Solutions
Biochemistry: Concepts and Connections (2nd Edition)
Ch. 8 - Prob. 1PCh. 8 - The enzyme urease catalyzes the hydrolysis of urea...Ch. 8 - An enzyme contains an active site aspartic acid...Ch. 8 - The folding and unfolding rate constants for a...Ch. 8 - In some reactions, in which a protein molecule is...Ch. 8 - Would you expect an “enzyme” designed to bind to...Ch. 8 - The initial rate for an enzyme-catalyzed reaction...Ch. 8 - a. If the total enzyme concentration in Problem 7...Ch. 8 - Prob. 9PCh. 8 - Prob. 10P
Ch. 8 - The following data describe the catalysis of...Ch. 8 - At 37 oC, the serine protease subtilisin has kcat...Ch. 8 - The accompanying figure shows three...Ch. 8 - The steady-state kinetics of an enzyme are studied...Ch. 8 - The same enzyme as in Problem 14 is studied in the...Ch. 8 - Enalapril is an anti-hypertension “pro-drug"...Ch. 8 - Initial rate data for an enzyme that obeys...Ch. 8 - Prob. 18PCh. 8 - Suggest the effects of each of the following...Ch. 8 - The inhibitory effect of an uncompetitive...Ch. 8 - Prob. 21PCh. 8 - Prob. 22PCh. 8 - Prob. 23PCh. 8 - In kinetics experiments, the hydrolysis of the...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biochemistry and related others by exploring similar questions and additional content below.Similar questions
- Fura-2 Fluorescence (Arbitrary Unit) 4500 4000 3500 3000 2500 2000 1500 1000 500 [Ca2+]=2970nM, 25°C [Ca2+] 2970nM, 4°C [Ca2+]=0.9nM, 25°C [Ca2+] = 0.9nM, 4°C 0 260 280 300 340 360 380 400 420 440 Wavelength (nm) ← < The figure on the LHS shows the excitation spectra of Fura-2 (Em = 510 nm) in 2 solutions with two different Ca2+ ion concentration as indicated. Except for temperature, the setting for excitation & signal acquisition was identical.< ப a) The unit in Y-axis is arbitrary (unspecified). Why? < < b) Compare & contrast the excitation wavelength of the Isosbestic Point of Fura-2 at 25 °C & 4 °C. Give a possible reason for the discrepancy. < c) The fluorescence intensity at 25 °C & 4 °C are different. Explain why with the concept of electronic configuration. <arrow_forwarddraw in the structure of each amino acid (as L-amino acids) using the Fischer projection style. an example has been included. Draw the structure for glycine, alanine, valine, isoleucine, methionine, proline, phenylalanine, tryptophan, serine, threonine, asparagine, glutamine, lysine, arginine, aspartic acid, glutamic acid, histidine, tyrosine, cysteinearrow_forwarddraw in the structure of each amino acid (as L-amino acids) using the Fischer projection style. an example has been includedarrow_forward
- draw in the structure of each amino acid (as L-amino acids) using the Fischer projection style. an example has been includedarrow_forwardDraw out the following peptide H-R-K-E-D at physiological pH (~7.4). Make sure toreference table 3.1 for pKa values.arrow_forwardThe table provides the standard reduction potential, E', for relevant half-cell reactions. Half-reaction E'° (V) Oxaloacetate² + 2H+ + 2e malate²- -0.166 Pyruvate + 2H+ + 2e → lactate -0.185 Acetaldehyde + 2H+ + 2e¯ →→→ ethanol -0.197 NAD+ + H+ + 2e--> NADH -0.320 NADP+ + H+ + 2e →→ NADPH Acetoacetate + 2H+ + 2e¯ - -0.324 B-hydroxybutyrate -0.346 Which of the reactions listed would proceed in the direction shown, under standard conditions, in the presence of the appropriate enzymes? Malate + NAD+ oxaloacetate + NADH + H+ Malate + pyruvate oxaloacetate + lactate Pyruvate + NADH + H+ lactate + NAD+ Pyruvate + p-hydroxybutyrate lactate + acetoacetate Acetaldehyde + succinate ethanol + fumerate Acetoacetate + NADH + H+ → B-hydroxybutyrate + NAD+arrow_forward
- Arrange the four structures in order from most reduced to most oxidized. Most reduced R-CH2-CH3 R-CH2-CH₂-OH R-CH,-CHO R-CH₂-COO Most oxidizedarrow_forwardfor each pair of biomolecules, identify the type of reaction (oxidation-reduction, hydrolysis, isomerization, group transfer, or nternal rearrangement) required to convert the first molecule to the second. In each case, indicate the general type of enzyme and cofactor(s) c reactants required, and any other products that would result. R-CH-CH-CH-C-S-COA A(n) A(n) A(n) A(n) Palmitoyl-CoA R-CH-CH=CH-C-S-CoA ° trans-A-Enoyl-CoA reaction converts palmitoyl-CoA to trans-A2-enoyl-CoA. This reaction requires and also produces Coo HN-C-H CH₂ CH₂ CH CH CH, CH, L-Leucine CH, CH, D-Leucine 8/6881 COO HÌNH: reaction converts L-leucine to D-leucine. This reaction is catalyzed by a(n) H-C-OH H-C-OH C=0 HO-C-H HO-C-H H-C-OH H-C-OH H-C-OH CH,OH Glucose H-C-OH CH,OH Fructose OH OH OH CH-C-CH₂ reaction converts glucose to fructose. This reaction is catalyzed by a(n) OH OH OPO I CH-C-CH H Glycerol Glycerol 3-phosphate H reaction converts glycerol to glycerol 3-phosphate. This reaction requires H,N- H,N H…arrow_forwardAfter adding a small amount of ATP labeled with radioactive phosphorus in the terminal position, [7-32P]ATP, to a yeast extract, a researcher finds about half of the 32P activity in P; within a few minutes, but the concentration of ATP remains unchanged. She then carries out the same experiment using ATP labeled with 32P in the central position, [ẞ-³2P]ATP, but the 32P does not appear in P; within such a short time. Which statements explain these results? Yeast cells reincorporate P; released from [ß-³2P]ATP into ATP more quickly than P¡ released from [y-³2P]ATP. Only the terminal (y) phosphorous atom acts as an electrophilic target for nucleophilic attack. The terminal (y) phosphoryl group undergoes a more rapid turnover than the central (B) phosphate group. Yeast cells maintain ATP levels by regulating the synthesis and breakdown of ATP. Correct Answerarrow_forward
- Compare the structure of the nucleoside triphosphate CTP with the structure of ATP. NH₂ 0- 0- 0- ·P—O—P—O—P—O—CH₂ H H H H OH OH Cytidine triphosphate (CTP) Consider the reaction: ATP + CDP ADP + CTP NH 0- 0- 0- ¯0— P—O— P—O—P-O-CH₂ H Η о H H OH OH Adenosine triphosphate (ATP) NH₂ Now predict the approximate K'eq for this reaction. Now predict the approximate AG for this reaction. Narrow_forwardThe standard free energy, AGO, of hydrolysis of inorganic polyphosphate, polyP, is about −20 kJ/mol for each P; released. In a cell, it takes about 50 kJ/mol of energy to synthesize ATP from ADP and Pi. ○ P O Inorganic polyphosphate (polyP) Is it feasible for a cell to use polyP to synthesize ATP from ADP? Why or why not? No. The reaction is unidirectional and always proceeds in the direction of polyP synthesis from ATP. Yes. If [ADP] and [polyP] are kept high, and [ATP] is kept low, the actual free-energy change would be negative. No. The synthesis of ATP from ADP and P; has a large positive G'o compared to polyP hydrolysis. Yes. The hydrolysis of polyP has a sufficiently negative AG to overcome the positive AGO of ATP synthesis. Correct Answerarrow_forwardIn the glycolytic pathway, a six-carbon sugar (fructose 1,6-bisphosphate) is cleaved to form two three-carbon sugars, which undergo further metabolism. In this pathway, an isomerization of glucose 6-phosphate to fructose 6-phosphate (as shown in the diagram) occurs two steps before the cleavage reaction. The intervening step is phosphorylation of fructose 6-phosphate to fructose 1,6-bisphosphate. H H | H-C-OH H-C-OH C=0 HO-C-H HO-C-H phosphohexose isomerase H-C-OH H-C-OH H-C-OH H-C-OH CH₂OPO CH₂OPO Glucose 6-phosphate Fructose 6-phosphate What does the isomerization step accomplish from a chemical perspective? Isomerization alters the molecular formula of the compound, allowing for subsequent phosphorylation. Isomerization moves the carbonyl group, setting up a cleavage between the central carbons. Isomerization causes the gain of electrons, allowing for the eventual release of NADH. Isomerization reactions cause the direct production of energy in the form of ATP.arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning Anatomy & PhysiologyBiologyISBN:9781938168130Author:Kelly A. Young, James A. Wise, Peter DeSaix, Dean H. Kruse, Brandon Poe, Eddie Johnson, Jody E. Johnson, Oksana Korol, J. Gordon Betts, Mark WomblePublisher:OpenStax College
Anatomy & PhysiologyBiologyISBN:9781938168130Author:Kelly A. Young, James A. Wise, Peter DeSaix, Dean H. Kruse, Brandon Poe, Eddie Johnson, Jody E. Johnson, Oksana Korol, J. Gordon Betts, Mark WomblePublisher:OpenStax College

Human Heredity: Principles and Issues (MindTap Co...
Biology
ISBN:9781305251052
Author:Michael Cummings
Publisher:Cengage Learning

Biology: The Dynamic Science (MindTap Course List)
Biology
ISBN:9781305389892
Author:Peter J. Russell, Paul E. Hertz, Beverly McMillan
Publisher:Cengage Learning

Biochemistry
Biochemistry
ISBN:9781305577206
Author:Reginald H. Garrett, Charles M. Grisham
Publisher:Cengage Learning


Biochemistry
Biochemistry
ISBN:9781305961135
Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougal
Publisher:Cengage Learning

Anatomy & Physiology
Biology
ISBN:9781938168130
Author:Kelly A. Young, James A. Wise, Peter DeSaix, Dean H. Kruse, Brandon Poe, Eddie Johnson, Jody E. Johnson, Oksana Korol, J. Gordon Betts, Mark Womble
Publisher:OpenStax College
Biomolecules - Protein - Amino acids; Author: Tutorials Point (India) Ltd.;https://www.youtube.com/watch?v=ySNVPDHJ0ek;License: Standard YouTube License, CC-BY