
Concept explainers
Interpretation:
The case of a protein that does not include methionine as its first amino acid has to be accounted.
Concept Introduction:
RNA synthesis: The process of RNA synthesis is Transcription. A small section of DNA unwinds, only one of the two strands act as template and the other strand as informational strand. The complementary bases are attached one by one by the action of RNA polymerase at template strand on moving down. The newly generated RNA is the exact copy of the informational strand, with the exception that a U replaces each T in the template DNA. The RNA synthesised carries genetic information and directs protein synthesis.
Codon: A sequence of three ribonucleotides in the mRNA chain that codes for a specific amino acid; also a three-
Genetic code: The sequence of nucleotides, coded in triplets (codons) in mRNA that determines the sequence of amino acids in protein synthesis.
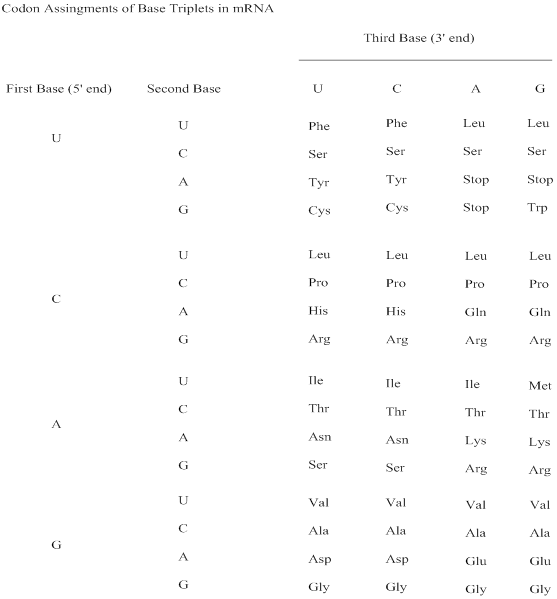
Illustrated relationships are:
DNA informational strand: 5’ ATG CCA GTA GGC CAC TTG TCA 3’
DNA Template strand: 3’ TAC GGT CAT CCG GTG AAC AGT 5’
mRNA: 5’ AUG CCA GUA GGC CAC UUG UCA 3’
protein: Met Pro Val Gly His Leu Ser
Notice: 5’ end of the mRNA strand codes for the N-terminal amino acid, whereas the 3’ end of the mRNA strand codes for the C-terminal amino acid. Proteins are always written N-terminal to C-terminal, reading left to right.
Want to see the full answer?
Check out a sample textbook solution
Chapter 26 Solutions
Fundamentals of General, Organic, and Biological Chemistry (8th Edition)
- Show the fate of the proton on the 4-Oxygen molecule of F-1,6-BP. Please include a drawing showing the electron flow that occurs.arrow_forward1. Which one is the major organic product obtained from the following aldol condensation? O NaOH, H₂O heat A B C D Earrow_forwardAn organic chemist ordered the wrong item. She wanted to obtain 1-hydroxy-2-butanone, butinstead ordered 2-hydroxybutyraldehyde. As a good biochemist, show how the organic chemistcould use biological catalysis to make her desired compound. Please show the mechanism by drawing.arrow_forward
- Show the fate of the hydrogen on carbon-2 of glucose. Please draw out the structure using curve arrows to show electron flow.arrow_forward3. Which one of the compounds below is the major product formed by the reaction sequence shown here? CH3 + CH3NO2 NaOH H2, Ni ? nitromethane acetophenone OH OH HO HN- u x x x x Ph A HO -NH2 HO H Ph Ph Ph N- H B Ph NH2 D Earrow_forward4. Only ONE of the five compounds below can be prepared by an aldol condensation in which a single carbonyl compound is treated with base. Which one is it? To solve this problem, reverse the aldol condensation that formed each of these molecules to find out what two molecules came together to make the products. The one in which the two molecules are identical is the answer. Ph Ph ཚིག གནས ག ནཱ ཀ ན ཀནཱ A Ph H B Ph Ph H D Ph. Ph Ph E Harrow_forward
- 5. Which one is the major organic product obtained from the following reaction sequence? First, equimolar amounts of cyclopentanone and LDA are mixed at -78°C. Then propionaldehyde (propanal) is added. Addition of aqueous acid completes the process. LDA, -78°C. 1. 2. H₂O* H A B H 0 D H H Earrow_forward2. Which one is the major organic product obtained from the following reaction? NaOH, H₂O heat A B C D Earrow_forwardCH3CH2CHO + propanal PhCH2CHO 2-phenylacetaldehyde mixture of four products NaOH 10. In the crossed aldol reaction of propanal and 2-phenylacetaldehyde shown above, a mixture of four products will be formed. Which ONE of the compounds below will NOT be formed in this crossed aldol reaction? OH Ph A H OH OH Ph H B OH OH H H H Ph Ph C Ph D Earrow_forward
- An organic chemist ordered the wrong item. She wanted to obtain 1-hydroxy-2-butanone, butinstead ordered 2-hydroxybutyraldehyde. As a good biochemist, show how the organic chemistcould use biological catalysis to make her desired compound.arrow_forwardPredict the products of aldolase catalyzing the reaction with acetone and (S)-3-hydroxybutyraldehyde.arrow_forwardA cancer patient undergoing chemotherapy is taking a protein kinase inhibitor drug. The patientis an avid marathon runner and does not want to miss his upcoming race. Is it a good idea forthis patient to attempt a marathon while on this medication? Explain why or why not.arrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning
Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning Human Biology (MindTap Course List)BiologyISBN:9781305112100Author:Cecie Starr, Beverly McMillanPublisher:Cengage Learning
Human Biology (MindTap Course List)BiologyISBN:9781305112100Author:Cecie Starr, Beverly McMillanPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning





