
Concept explainers
Northern blot analysis is performed on mRNA produced by transcription of a gene in organisms with different genotypes. Three alleles occur at the gene: N is a dominant wild
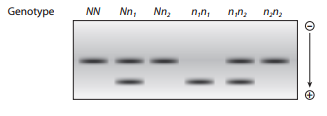
Organisms with the genotypes NN,
Organisms that are n1n1 have a single mRNA band with higher electrophoretic mobility. Thinking about the composition of mRNA, explain this observation.
Two mRNA bands are detected for organisms with the
Want to see the full answer?
Check out a sample textbook solution
Chapter 10 Solutions
Genetic Analysis: An Integrated Approach (2nd Edition)
- The following table lists 4 bacterial strains that are partial diploids for lac operon genes. Given the activity of beta-galactosidase measured for each strain in the absence (-lac) or presence (+lac) of lactose, complete the table by choosing the appropriate symbol (+, -, C, S) to indicate the allele of the gene or site missing from the table (blue numbers). strain A BC 5 C D 7 chromosome I O 1 2 4 1 [Select] 9 3 [Select] [Select] [Select] 9 [Select] + + Z + + 6 + I +5 + 10 plasmid O 3 + 7 C Z + 8 8 2 [Select] 4 [Select] 6 [Select] B-gal act. -lac +lac 0.002 0.003 0.002 0.058 0.063 0.121 0.059 0.062 Select] 1 ✔ [ Select] + is C Sarrow_forwardA complex biochemical pathway is shown below, along with the alleles that either promote or inhibit each step of the pathway leading to a phenotype. Gene A has alleles A and a, B has alleles B and b, and so forth. Genes B and C are duplicate dominant epistatic lethal as heterozygotes (i.e. Bb Cc are lethal). Genes D and E are duplicate dominant epistatic (i.e. dd eg = desired phenotype). If I were to cross AA Bb cc Dd Ee with aa BB Cc Dd e, (i) (ii) What proportion of all offspring don't show the phenotype? What proportion of offspring survive? Gene A Gene B B Gene D a Gene C Gene Earrow_forwardWhen considering the cause of the disease in a patient who is the child of an inbred couple, it was found that 4 enzymes in the lysosome were inactivated. When comparing the amino acid sequences of these four enzymes with the wild-type enzyme, no difference was found in the number and arrangement of amino acids. Given mutations that may be the cause of the patient's disease above.arrow_forward
- 1) Using the diagram below, sketch in the pattern of bands you would expect to see after digesting the DNA of the TT, Tt and tt genotypes of the TAS2R38 gene. Use the Base Pair Standards on the left of the diagram as a reference in drawing the positions of the bands. Base Pair TASTER GENO TYPE Standards (Base Pairs) TT Tt tt 500 400 300 200 100 50arrow_forwardA yeast geneticist irradiates haploid cells of a strain thatis an adenine-requiring auxotrophic mutant, caused bymutation of the gene ade1. Millions of the irradiatedcells are plated on minimal medium, and a small number of cells divide and produce prototrophic colonies.These colonies are crossed individually with a wildtype strain. Two types of results are obtained:(1) prototroph × wild type : progeny all prototrophic(2) prototroph × wild type : progeny 75% prototrophic,25% adenine-requiring auxotrophsa. Explain the difference between these two types ofresults.b. Write the genotypes of the prototrophs in each case.c. What progeny phenotypes and ratios do you predictfrom crossing a prototroph of type 2 by the original ade1auxotroph?arrow_forwardIn E. coli, four Hfr strains donate the following markers,shown in the order donated:Strain 1: M Z X W CStrain 2: L A N C WStrain 3: A L B R UStrain 4: Z M U R BAll these Hfr strains are derived from the same F+ strain.What is the order of these markers on the circularchromosome of the original F+?arrow_forward
- Cystic fibrosis (CF) is a genetic disorder affecting a number of organs, including the lung airways, pancreas, and sweat glands. Mutations in both copies of the CFTR gene causes cystic fibrosis. Imagine that you have sweat gland samples from several Cystic Fibrosis patients (A-C) with unknown mutations in CFTR. You also have normal (+) sweat gland sample to use as a positive control. А В С А В С Choose which mutation would explain the RNA and protein results in A, B, & C: 1. Promoter/Regulatory mutation 2. Silent mutation 3. Missense mutation 4. Deletion mutation 5. Splice site mutation 6. Nonsense mutation RNA gel Protein gelarrow_forwardThe wildtype sequence of a gene is the following: wt: 3' TAC AAA TCT AGC CCG 5' and the following 3 mutations were found:, indicate what type of mutation this is 1: 3' TAC AAA TCA AGC CCG 5' ; 2. 3' TAC AAA TCT ATC CCG 5'. ; 3. 3' TAC AAA AGC CCG 5'. ;arrow_forwardThe DNA sequence of one strand of a gene from threeindependently isolated mutants is given here (5′ endsare at left). Using this information, what is the sequence of the wild-type gene in this region?mutant 1 ACCGTAATCGACTGGTAAACTTTGCGCGmutant 2 ACCGTAGTCGACCGGTAAACTTTGCGCGmutant 3 ACCGTAGTCGACTGGTTAACTTTGCGCGarrow_forward
- In a transformation experiment involving a wild type bacterial strain with a recipient strain with mutations in genes f, g, h and i pairs of genes were analyzed for co-transformation with the following results: Gene Pair Co-transformation g+ i+ f+ i+ yes no f+ h+ Уes f+ g+ no g+ h+ h+ i+ no уes What is the linear order of these genes relative to cach other?arrow_forwardGenomic DNA from a family where sickle-cell disease is known to be hereditary, is digested with the restriction enzyme MstII and run in a Southern Blot. The blot is hybridised with two different 0.6 kb probes, both probes (indicated in red in the diagram below) are specific for the β-globin gene (indicated as grey arrow on the diagram below). The normal wild-type βA allele contains an MstII restriction site indicated with the asterisk (*) in the diagram below; in the mutated sickle-cell βS allele this restriction site has been lost. What size bands would you expect to see on the Southern blots using probe 1 and probe 2 for an individual with sickle cell disease (have 2 βS alleles)? Probe 1 Probe 2 (a) 0.6kb 0.6kb and 1.2kb (b) 0.6kb and 1.8kb 0.6kb, 1.2kb and 1.8kb (c) 1.2kb 0.6kb (d) 1.8kb 1.8kb a. (a) b. (b) c. (c) d. (d)arrow_forwardThe following recombinants are recovered when conjugation occurs between an a*d*g+ donor and an adg recipient. at d+ g+ = 84% a d g+ = 6% at d g+ = 10% a dt g+ = less than 1% What is the map distance between the a and d genes? 10 map units 74 map units less than 1 map unit 84 map units 6 map unitsarrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





