
Concept explainers
An Hfr strain with the genotype cys+leu+met+strS is mated with an F- strain carrying the genotype cys- leu- met- strR. In an interrupted mating experiment, small samples of the conjugating bacteria are withdrawn every
Medium
Medium
Medium
a. What donor gene is the selected marker in each medium?
b. List all possible bacterial genotypes growing on each medium.
c. What is the purpose of adding streptomycin to each selection medium? The table on next page shows the number of colonies growing on each selection medium. The sampling time indicates how many minutes have passed since conjugation began
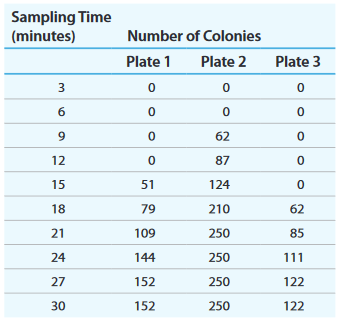
d. Determine the order of donor genes cys, leu, and met from the interrupted mating data.
e. Suppose a fourth selection medium containing leucine and streptomycin is prepared. At what sampling time do you expect the first-growing colonies to appear? Explain your reasoning.
Want to see the full answer?
Check out a sample textbook solution
Chapter 6 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- Each of the three different Hfr strains in the table below (A, B and C) arose independently and possess the same markers. Each of the three Hfr strains was separately mixed with the same F strain carrying the alternative markers. Conjugation was interrupted at various times after mixing over a period of 95 minutes. The bacteria were then plated on the appropriate media and recipients scored for various Hfr markers. The times (in minutes after mixing) at which recombinants carrying particular donor markers first appear, and therefore the times at which these first enter the F- cell and then integrate into the recipient genome, are shown below: Donor markers Time of appearance in minutes Strain A Strain B Strain C his+ 6.5 68.5 29.5 gal+ 30.0 46.0 52.0 pro+ 34.5 41.5 56.5 met+ 58.0 20.0 76.0 mtl+ 67.0 11.0 85.0 xyl+ 68.5 9.5 86.5 mal+ 73.0 5.0 3.0 ser+ 81.5 86.5 11.5 tyr+ 88.0 78.5 19.5 Draw the genome of each strain showing the location of the markers, the location of F, and the…arrow_forwardYou co-culture the following bacterial strains: an Hfr prototroph and an F- auxotroph for the genes mal, met, mtl, and xyl. You interrupt conjugation at various time points and place the mixtures on media plates lacking each of the nutrients. Based on the results shown on the right, what is the order of these four genes along the bacterial chromosome? Explain your reasoning.arrow_forwardDNA from a strain of Bacillus subtilis with genotype a+ b+ c+ d+ e+ is used to transform a strain with genotype a− b− c− d− e−. Pairs of genes are checked for cotransformation, and the following results are obtained: Pair of genes Cotransformation Pair of genes Cotransformation a+ and b+ No b+ and d+ No a+ and c+ No b+ and e+ Yes a+ and d+ Yes c+ and d+ No a+ and e+ Yes c+ and e+ Yes b+ and c+ Yes d+ and e+ No On the basis of these results, what is the order of the genes on the bacterial chromosome?arrow_forward
- You co-culture the following bacterial strains: an Hfr prototroph and an F- auxotroph for the genes mal, met, mtl, and xyl. You interrupt conjugation at various time points and place the mixtures on media plates lacking each of the nutrients. Based on the results shown in the image, what is the order of these four genes along the bacterial chromosome? Please explain!arrow_forwardUsing bacteriophage P22, you performed a threefactor cross in Salmonella typhimurium. The crosswas between an Arg− Leu− His− recipient bacteriumand bacteriophage P22, which was grown on an Arg+Leu+ and His+ strain. You selected for 1000 Arg+transductants and tested them on several selective media by replica plating. You obtained the followingresults:Arg+ Leu− His− 585Arg+ Leu− His+ 300Arg+ Leu+ His+ 114Arg+ Leu+ His− 1a. What are the cotransduction frequencies of argwith either his or leu?b. Is arg closer to his than it is to leu, or is arg closerto leu than it is to his? What are the relative mapdistances?c. Why can you not obtain an accurate cotransductionfrequency for his and leu from the data provided?d. What is the order of the three markers?e. Draw the crossovers that would produce the classwith only one transductant.f. Estimate the proportion of the 4.9 Mb S. typhimuriumgenome that contains these three genes. Assume thatbacteriophage P22 can package the same amountof DNA as the…arrow_forwardFour Hfr strains are derived from an F+ strain of E. Coli to serve as donors for an interrupted-mating experiment. Use the time-table and partial map of the F+ strain (shown below) to determine the genes’ respective positions. Keep in mind that the map distances are NOT proportional, only the FIRST 5 markers are indicated per strain, and the entry times, recorded in minutes, are in parentheses. Transferred genes represent wild-type alleles. Based on the data, which gene can be located at position 5 on the map?arrow_forward
- DNA obtalned from the indicated donor strains is used to TRANSFORM the indicated recipient strains. The resulting progeny are plated on minimal medium so that only wild- type recombinants are scored. The number of wild-types for each cross is given in the chart below. What is the order of the genes? Donor Recipient wid type colonies a-b- c+ a+ b+ c- 273 a- b+ c- a+ b-c+ 462 a- b+ C+ a+ b-c- 2 a b- C+ a- b+ c+ O 4-b-c a-o-b b-a-c cannot be determined Question 18 of 26arrow_forwardThree different Hfr donor strains are mixed with separate samples of an F− strain, and the following mapping data are provided from studies of interrupted conjugation: Appearance of genes in F − cells Hfr1: Genes b+ d+ c+ f+ g+ Time* 3 5 16 27 59 Hfr2: Genes e+ f+ c+ d+ b+ Time 6 24 35 46 48 Hfr3: Genes d+ c+ f+ e+ g+ Time 4 15 26 44 58 Construct a genetic map for these genes, indicating their order on the bacterial chromosome and the distances between them.arrow_forward5 strains of E. coli are isolated. They may be mutated on the ara, mal, leu and pro genes, and only these 4 genes. The 5 strains (E1 to E5) are tested on 5 different media and colonies are observed the next day. Rich Media (Lb) E5 E4 E1 E2Ⓡ E3 MM + Mal+Leu E5 E4 X ● E1 E5 X MM + Mal E4 X X E1 E3 E2 E5 ✪✪ E2 E4 ● E1 E5 MM + Mal+Leu +Pro E2 E3 MM + Ara E4 X X E1 E3 E2 X X Colonies are indicated by a black dot, absence of a colony is indicated by a cross. 1) Determine the genotypes of the 5 bacterial strains (E1 - E5)arrow_forward
- A prototrophic strain (his" arg" lac') was used as a donor to transform an auxotrophic strain (his arg lac'). Initial transformants are isolated on minimal medium + histidine + arginine + lactose - glucose. i. What genotypes will grow on this medium? ii. These colonies are replicated to minimal medium + histidine, and 50% of the original colonies grow. What genotypes will grow on this medium? iii. The original colonies are also replicated to minimal medium + arginine, and 10% of the colonies grow. What genotypes will grow on this medium? iv. The original colonies are also replicated to minimal medium. No colonies grow. Based on this information, what genotypes will grow on minimal medium + histidine and on minimal medium + arginine? What is the relative gene order for his, arg, and lac? Which two genes are closer? Explain your answer.arrow_forwardWhen demonstrating horizontal gene transfer, why is it important to use strains that each require at least two different amino acids?arrow_forwardGiven what we've discussed in class, what will be most likely outcome if you conjugate an streptomycin resistant ampicillin sensitive methionine auxotroph E. coli strain (engineered to be pir+) that is F- with a streptomycin sensitive non-HFR methionine prototroph strain that is F- and RP4+ but contains pUC18? Colonies on minimal media + ampicillin +streptomycin plates No colonies on minimal media +ampicillin +streptomycin platesarrow_forward
 Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning

