
Study Guide for Campbell Biology
11th Edition
ISBN: 9780134443775
Author: Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Jane B. Reece, Martha R. Taylor, Michael A. Pollock
Publisher: PEARSON
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 20, Problem 6TYK
The following segment of DNA has restriction sites I and II, which create restriction fragments a, b, and c. Which of the gels produced by electrophoresis would represent the separation and identity of these fragments?
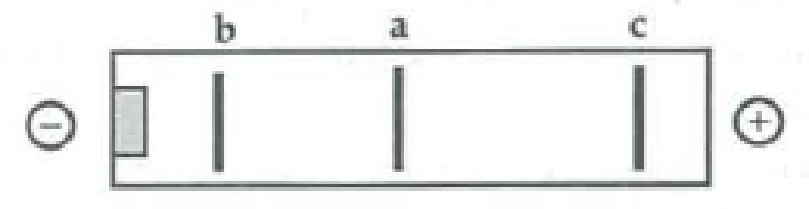

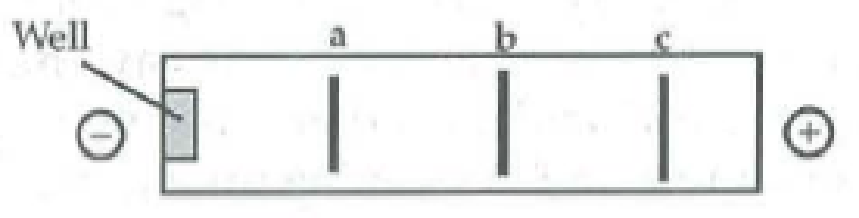


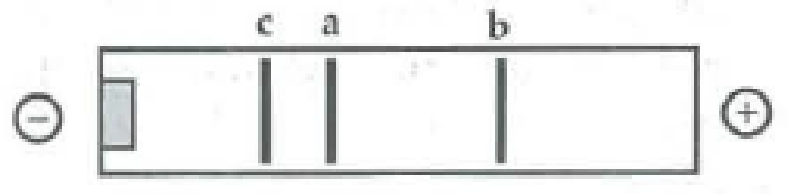
Expert Solution & Answer
Trending nowThis is a popular solution!

Students have asked these similar questions
As a medical professional, it is important to be able to discuss how genetic processes such as translation regulation can directly affect patients. Think about some situations that might involve translation regulation.
Respond to the following in a minimum of 175 words:
Why is translation regulation important?
What are some examples of translation regulation in humans?
Select one of the examples you provided and explain what happens when translation regulation goes wrong.
The metabolic pathway below is used for the production of the purine nucleotides adenosine monophosphate (AMP) and guanosine monophosphate (GMP) in eukaryotic cells. Assume each arrow represents a reaction catalyzed by a different enzyme. Using the principles of feedback inhibition, propose a regulatory scheme for this pathway that ensures an adequate supply of both AMP and GMP, and prevents the buildup of Intermediates A through G when supplies of both AMP and GMP are adequate.
QUESTION 27
Label the structures marked A, B, C and explain the role of structure A.
W
plasma membrane
For the toolbar, press ALT+F10 (PC) or ALT+FN+F10 (Mac).
BIUS
☐
Paragraph
Π " ΩΘΗ
Β
Open Sans, a...
10pt
EE
Chapter 20 Solutions
Study Guide for Campbell Biology
Ch. 20 - In what ways would third-generation sequencing be...Ch. 20 - The following schematic diagram depicts an...Ch. 20 - Which of the following DNA sequences would most...Ch. 20 - a. When PCR is used to prepare a DNA fragment for...Ch. 20 - a. What are some of the benefits of determining...Ch. 20 - Prob. 6IQCh. 20 - What are some of the practical and ethical...Ch. 20 - Prob. 8IQCh. 20 - Prob. 1SYKCh. 20 - Fill in the table on the previous page on the...
Ch. 20 - Prob. 3SYKCh. 20 - Prob. 1TYKCh. 20 - Prob. 2TYKCh. 20 - Gel electrophoresis is a means of separating...Ch. 20 - Prob. 4TYKCh. 20 - Prob. 5TYKCh. 20 - The following segment of DNA has restriction sites...Ch. 20 - Prob. 7TYKCh. 20 - Prob. 8TYKCh. 20 - Prob. 9TYKCh. 20 - Prob. 10TYKCh. 20 - Prob. 11TYKCh. 20 - Which enzyme is used in the polymerase chain...Ch. 20 - Prob. 13TYKCh. 20 - STRs (short tandem repeats) are a valuable tool...Ch. 20 - Prob. 15TYKCh. 20 - Which of the following has the greatest potential...Ch. 20 - Prob. 17TYKCh. 20 - Petroleum-lysing bacteria are being engineered for...Ch. 20 - Prob. 19TYKCh. 20 - Prob. 20TYK
Additional Science Textbook Solutions
Find more solutions based on key concepts
To test your knowledge, discuss the following topics with a study partner or in writing ideally from memory. Th...
HUMAN ANATOMY
Describe the role and impact of microbes on the earth.
Microbiology Fundamentals: A Clinical Approach
On what molecule does the anticodon appear? Explain the role of this molecule in protein synthesis.
Human Physiology: An Integrated Approach (8th Edition)
2. Which of the following is the best example of the use of a referent? _
a. A red bicycle
b. Big as a dump tru...
Physical Science
What process causes the Mediterranean intermediate Water MIW to become more dense than water in the adjacent At...
Applications and Investigations in Earth Science (9th Edition)
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- examples of synamptomorphyarrow_forwardexamples of synamtomorphy.arrow_forwardE. Bar Graph Use the same technique to upload the completed image. We will use a different type of graph to derive additional information from the CO2 data (Fig A1.6.2) 1. Calculate the average rate of increase in COz concentration per year for the time intervals 1959-1969, 1969- 1979, etc. and write the results in the spaces provided. The value for 1959-1969 is provided for you as an example. 2. Plot the results as a bar graph. The 1959-1969 is plotted for you. 3. Choose the graph that looks the most like yours A) E BAR GRAPH We will use a different type of graph to derive additional information from the CU, data (rig. nive). Average Yearly Rate of Observatory, Hawall interval Rate of increase per year 1959-1969 0.9 1969-1979 1979-1989 1989-1999 1999-2009 Figure A1.6.2 1999-2009 *- mrame -11- -n4 P2 جية 1989-1999 1979-1989 1969-1979 1959-1969 This bar drawn for you as an example 1.0 CO, Average Increase/Year (ppmv) B) E BAR GRAPH We will use a different type of graph to derive…arrow_forward
- Use the relationships you just described to compute the values needed to fill in the blanks in the table in Fig A1.4.1 depth (a) 1.0 cml 0.7 cml cm| base dimensions (b, c)| 1.0 cm| 1.0 cm| 1.0 cm 1.0 cm| 1.0 cm| 1.0 cm volume (V) 1.0_cm' cm'| cm'| density (p) 1.0 g/cm'| 1.0 g/cm 1.0 g/cm' mass (m)| 0.3 g Column 1: depth at 1.0 cm volume mass Column 2: depth at 0.7 cm volume mass Column 3: unknown depth depth volumearrow_forwardSan Andreas Transform Boundary Plate Motion The geologic map below of southern California shows the position of the famous San Andreas Fault, a transform plate boundary between the North American Plate (east side) and the Pacific Plate (west side). The relative motion between the plates is indicated by the half arrows along the transform plate boundary (i.e., the Pacific Plate is moving to the northwest relative to the North American Plate). Note the two bodies of Oligocene volcanic rocks (labeled Ov) on the map in the previous page located along either side of the San Andreas Fault. These rocks are about 23.5 million years old and were once one body of rock. They have been separated by displacement along the fault. 21. Based on the offset of these volcanic rocks, what is the average annual rate of relative plate motion in cm/yr? SAF lab 2.jpg Group of answer choices 0.67 cm/yr 2 cm/yr 6.7 cm/yr 1.5 cm/yr CALIFORNIA Berkeley San Francisco K Os Q San Andreas Fault Ov…arrow_forwardThese are NOT part of any graded assignment. Are there other examples of synapomorphy. What is it called when the traits retained are similar to ancestors?arrow_forward
- Please hand draw everying. Thank you! Draw a gram positive bacterial cell below. Your cell should have the following parts, labeled: A coccus shape A capsule The gram positive cell wall should have the peptidoglycan labeled, as well as its component parts (NAM, NAG, and teichoic acid) A cell membrane Fimbriae A nucleoid Ribosomes Inclusionsarrow_forwardDraw a gram negative bacterial cell below. Your cell should have the following parts, labeled: A bacillus shape Fimbriae Amphitrichous flagella 2 membranes (outer and inner) The outer membrane should have lipopolysaccharide (LPS) with lipid A and O antigens Periplasmic space The thin peptidoglycan cell wall between the 2 membranes A nucleoid Ribosomes Inclusionsarrow_forwardBacterial species Cell wall type Example: S. mitis Gram positive S. epidermidis H. pylori M. bovis S. marcescens Shape and arrangement Coccus, streptococcus Drawing 0000000arrow_forward
- Draw a gram positive bacterial cell below. Your cell should have the following parts, labeled: A coccus shape A capsule The gram positive cell wall should have the peptidoglycan labeled, as well as its component parts (NAM, NAG, and teichoic acid) A cell membrane Fimbriae A nucleoid Ribosomes Inclusionsarrow_forwardwhat rank is above kingdom? order, class, phylum or domainarrow_forwardin the hierarchy of taconomic categories, with kingdom at the top, what taxon is below classarrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning
Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning Human Biology (MindTap Course List)BiologyISBN:9781305112100Author:Cecie Starr, Beverly McMillanPublisher:Cengage Learning
Human Biology (MindTap Course List)BiologyISBN:9781305112100Author:Cecie Starr, Beverly McMillanPublisher:Cengage Learning Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning Concepts of BiologyBiologyISBN:9781938168116Author:Samantha Fowler, Rebecca Roush, James WisePublisher:OpenStax College
Concepts of BiologyBiologyISBN:9781938168116Author:Samantha Fowler, Rebecca Roush, James WisePublisher:OpenStax College Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax

Biology Today and Tomorrow without Physiology (Mi...
Biology
ISBN:9781305117396
Author:Cecie Starr, Christine Evers, Lisa Starr
Publisher:Cengage Learning

Human Biology (MindTap Course List)
Biology
ISBN:9781305112100
Author:Cecie Starr, Beverly McMillan
Publisher:Cengage Learning

Human Heredity: Principles and Issues (MindTap Co...
Biology
ISBN:9781305251052
Author:Michael Cummings
Publisher:Cengage Learning

Concepts of Biology
Biology
ISBN:9781938168116
Author:Samantha Fowler, Rebecca Roush, James Wise
Publisher:OpenStax College

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax

QCE Biology: Introduction to Gene Expression; Author: Atomi;https://www.youtube.com/watch?v=a7hydUtCIJk;License: Standard YouTube License, CC-BY