
Concepts of Genetics (12th Edition)
12th Edition
ISBN: 9780134604718
Author: William S. Klug, Michael R. Cummings, Charlotte A. Spencer, Michael A. Palladino, Darrell Killian
Publisher: PEARSON
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 6, Problem 16PDQ
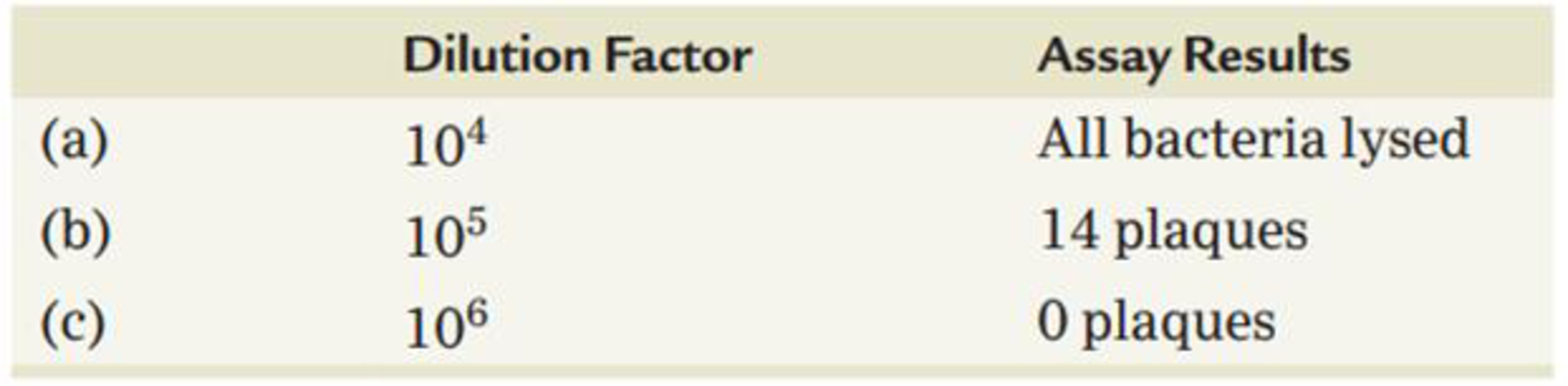
A phage-infected bacterial culture was subjected to a series of dilutions, and a plaque assay was performed in each case, with the results shown in the following table. What conclusion can be drawn in the case of each dilution, assuming that 0.1 mL was used in each plaque assay?
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
Case Study—Ella
Ella has a family history of diabetes. She wants to follow a healthful eating pattern that can lower her risk for developing this condition. Her dietitian recommends a goal of 450 to 600 kcal per meal and advises Ella to follow the Acceptable Macronutrient Distribution Range (AMDR) for carbohydrates and the Dietary Guidelines for Americans 2015-2020, which recommend limiting added sugar. She also recommends that Ella choose whole grains rather than processed grains. Ella decides to pack a lunch to take to work every day. This morning she’s making a sandwich for her lunch.
Categories of Sandwich Options (Top of the screen)
Breads
Spreads
Cheeses
Vegetables
Proteins
Specific food items to select
White Bread 6-inches
Honey Mustard
Provolone
LettuceTomatoBell Peppers
Turkey
Part A - Reading Nutrition Facts Panels for Total Kilocalories
How many total kilocalories are in Ella’s sandwich?
_______ kcal ?
Part B - Reading Nutrition Facts Panels for…
Case Study—Ella
Ella has a family history of diabetes. She wants to follow a healthful eating pattern that can lower her risk for developing this condition. Her dietitian recommends a goal of 450 to 600 kcal per meal and advises Ella to follow the Acceptable Macronutrient Distribution Range (AMDR) for carbohydrates and the Dietary Guidelines for Americans 2015-2020, which recommend limiting added sugar. She also recommends that Ella choose whole grains rather than processed grains. Ella decides to pack a lunch to take to work every day. This morning she’s making a sandwich for her lunch.
Categories of Sandwich Options (Top of the screen)
Breads
Spreads
Cheeses
Vegetables
Proteins
Specific food items to select
White Bread 6-inches
Honey Mustard
Provolone
LettuceTomatoBell Peppers
Turkey
Part A - Reading Nutrition Facts Panels for Total Kilocalories
How many total kilocalories are in Ella’s sandwich exactl ______kcal ?
Part B - Reading Nutrition Facts Panels for…
In humans, red-green color blindness is recessive and X-linked, whereas albinism is recessive and autosomal. What types of children can be produced as the result of marriage between two homozygous parents, a normal-vision albino woman and a color-blind, normal male?
Chapter 6 Solutions
Concepts of Genetics (12th Edition)
Ch. 6 - When the interrupted mating technique was used...Ch. 6 - In a transformation experiment involving a...Ch. 6 - In complementation studies of the rII locus of...Ch. 6 - A 4-month-old infant had been running a moderate...Ch. 6 - Prob. 2CSCh. 6 - Prob. 3CSCh. 6 - Prob. 4CSCh. 6 - HOW DO WE KNOW? In this chapter, we have focused...Ch. 6 - Review the Chapter Concepts list on p. 123. Many...Ch. 6 - With respect to F+ and F bacterial matings, answer...
Ch. 6 - List all major differences between (a) the F+ F...Ch. 6 - Describe the basis for chromosome mapping in the...Ch. 6 - In general, when recombination experiments are...Ch. 6 - Why are the recombinants produced from an Hfr F...Ch. 6 - Describe the origin of F bacteria and merozygotes.Ch. 6 - In a transformation experiment, donor DNA was...Ch. 6 - Describe the role of heteroduplex formation during...Ch. 6 - Explain the observations that led Zinder and...Ch. 6 - Prob. 12PDQCh. 6 - Two theoretical genetic strains of a virus (abc...Ch. 6 - The bacteriophage genome consists of many genes...Ch. 6 - If a single bacteriophage infects one E. coli cell...Ch. 6 - A phage-infected bacterial culture was subjected...Ch. 6 - In recombination studies of the rII locus in phage...Ch. 6 - In an analysis of rII mutants, complementation...Ch. 6 - If further testing of the mutations in Problem 18...Ch. 6 - Using mutants 2 and 3 from Problem 19, following...Ch. 6 - During the analysis of seven rII mutations in...Ch. 6 - In studies of recombination between mutants 1 and...Ch. 6 - Prob. 23ESPCh. 6 - An Hfr strain is used to map three genes in an...Ch. 6 - A plaque assay is performed beginning with 1 mL of...Ch. 6 - In a cotransformation experiment, using various...Ch. 6 - For the experiment in Problem 26, another gene, g,...Ch. 6 - Bacterial conjugation, mediated mainly by...Ch. 6 - A study was conducted in an attempt to determine...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- In Drosophila, an X linked recessive mutation, scalloped (sd), causes irregular wing margins. Diagram the F1 and F2 results if a (a) scalloped female is crossed with a normal male; (b) a scalloped male is crossed with a normal female (assume the female is homozygous). Compare these results to what you would find if the trait was not sex linked.arrow_forwardCase Study—Ella Ella has a family history of diabetes. She wants to follow a healthful eating pattern that can lower her risk for developing this condition. Her dietitian recommends a goal of 450 to 600 kcal per meal and advises Ella to follow the Acceptable Macronutrient Distribution Range (AMDR) for carbohydrates and the Dietary Guidelines for Americans 2015-2020, which recommend limiting added sugar. She also recommends that Ella choose whole grains rather than processed grains. Ella decides to pack a lunch to take to work every day. This morning she’s making a sandwich for her lunch. Categories of Sandwich Options (Top of the screen) Breads Spreads Cheeses Vegetables Proteins Specific food items to select White Bread 6-inches Honey Mustard Provolone LettuceTomatoBell Peppers Turkey Part A - Reading Nutrition Facts Panels for Total Kilocalories How many total kilocalories are in Ella’s sandwich exactl ______kcal ? Part B - Reading Nutrition Facts Panels for…arrow_forwardC MasteringHealth MasteringNu × session.healthandnutrition-mastering.pearson.com/myct/itemView?assignment ProblemID=17465255&attemptNo=1&offset=prevarrow_forwardBiopharmaceutics and Pharmacokinetics:Two-Compartment Model Instant Absorption Questions SHOW ALL WORK, including equation used, variables used and each step to your solution, report your regression lines and axes names (with units if appropriate) :Calculate a-q a) B1, b) B2, c) hybrid rate constant (1) d) hybrid rate constant (2) e) t1/2,dist f) t1/2,elim g) k10 h) k12 i) k21 j) initial concentration (C0) k) central compartment volume (V1) l) steady-state volume (Vss) m) clearance (CL) AUC (0→10 min) using trapezoidal rule n) AUC (20→30 min) using trapezoidal rule o) AUCtail (AUC360→∞) p) total AUC (using short cut method) q) volume from AUC (VAUC)arrow_forwardGlitazones reduce insulin resistance by binding to a transcription factor in adipocytes, thereby reducing thesecretion of fatty acids. Glitazones are taken orally (in pill form). Using pharmacokinetic modeling, deriveequations to describe how the concentration of glitazones varies in the plasma as a function of time. Yourequations should be of the form: dCglitazone /dt = something, or dMglitazone /dt = something. Your model shouldinclude three compartments: the gut, the plasma, and the fatty tissues. Make sure to include a diagram thatillustrates your thinking, state all assumptions, and define your variables. Do not solve the equations.arrow_forwardCase Study—Ella Review the case study and then answer Parts A through F. Ella has a family history of diabetes. She wants to follow a healthful eating pattern that can lower her risk for developing this condition. Her dietitian recommends a goal of 450 to 600 kcal per meal and advises Ella to follow the Acceptable Macronutrient Distribution Range (AMDR) for carbohydrates and the Dietary Guidelines for Americans 2015-2020, which recommend limiting added sugar. She also recommends that Ella choose whole grains rather than processed grains. Ella decides to pack a lunch to take to work every day. This morning she’s making a sandwich for her lunch. Categories of Sandwich Options (Top of the screen) Breads Spreads Cheeses Vegetables Proteins Specific food items to select White Bread 6-inches Honey Mustard Provolone LettuceTomatoBell Peppers Turkey Part A - Reading Nutrition Facts Panels for Total Kilocalories How many total kilocalories are in Ella’s sandwich? _____…arrow_forward, if one of the archaeological specimens lacked the celiac disease-causing epitope, how could PCR be used to identify the allele in a contemporary germplasm collection of wild wheats, and to assist in transferring the allele to modern wheat varieties?arrow_forwardNow you will consider the composition of lipoproteins, including where they are synthesized, how they circulate, and where the various lipid and protein components are located within the lipoprotein molecule. Drag the appropriate labels to their respective targets.arrow_forwardThe Oregon Wolfe Barley mapping population is unique in having 12 easily-scored morphological markers, each showing monogenic inheritance. Do you consider these markers useful? Briefly defend your answer, pointing out advantages and disadvantages of morphological vs. molecular markers.arrow_forwardBiopharmaceutics and Pharmacokinetics:Two-Compartment Model Instant Absorption Questions Calculate these : a) B1, b) B2, c) hybrid rate constant (1) d) hybrid rate constant (2) e) t1/2,dist f) t1/2,elim g) k10 h) k12 i) k21 j) initial concentration (C0) k) central compartment volume (V1) l) steady-state volume (Vss) m) clearance (CL) AUC (0→10 min) using trapezoidal rule n) AUC (20→30 min) using trapezoidal rule o) AUCtail (AUC360→∞) p) total AUC (using short cut method) q) volume from AUC (VAUC)arrow_forwardIn a population of Jackalopes (pictured below), horn length will vary between 0.5 and 2 feet, with the mean length somewhere around 1.05 feet. You pick Jackalopes that have horn lengths around 1.75 feet to breed as this appears to be the optimal length for battling other Jackalopes for food. After a round of breeding, you measure the offsprings' mean horn length is 1.67. What is the heritability of horns length (h2)? Is Jackalope horn length a heritable trait? (4 pts)? 12pt v R Paragraph V BIU A श्र > Barrow_forwardThere are many differences between DNA replication happening during mitosis in a Douglas fir tree growing in the Oregon Cascade Mountains and DNA replication happening during a PCR reaction in a forestry research lab at Oregon State University where the laboratory is amplifying a Simple Sequence Repeat. Complete the following table that compares the two DNA replication events in terms of the primers, the nucleotides, the polymerase, and the target sequence. Additionally, give a general value for the number of copies of the template DNA after one S phase in one cell and after the lab has completed the PCR reaction. Tree SSR Type your answer here: Primers Nucleotides Polymerase Target sequence Number of copiesarrow_forwardarrow_back_iosSEE MORE QUESTIONSarrow_forward_ios
Recommended textbooks for you
 Principles Of Radiographic Imaging: An Art And A ...Health & NutritionISBN:9781337711067Author:Richard R. Carlton, Arlene M. Adler, Vesna BalacPublisher:Cengage Learning
Principles Of Radiographic Imaging: An Art And A ...Health & NutritionISBN:9781337711067Author:Richard R. Carlton, Arlene M. Adler, Vesna BalacPublisher:Cengage Learning Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage LearningSurgical Tech For Surgical Tech Pos CareHealth & NutritionISBN:9781337648868Author:AssociationPublisher:CengageEssentials of Pharmacology for Health ProfessionsNursingISBN:9781305441620Author:WOODROWPublisher:Cengage
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage LearningSurgical Tech For Surgical Tech Pos CareHealth & NutritionISBN:9781337648868Author:AssociationPublisher:CengageEssentials of Pharmacology for Health ProfessionsNursingISBN:9781305441620Author:WOODROWPublisher:Cengage



Principles Of Radiographic Imaging: An Art And A ...
Health & Nutrition
ISBN:9781337711067
Author:Richard R. Carlton, Arlene M. Adler, Vesna Balac
Publisher:Cengage Learning

Biology: The Dynamic Science (MindTap Course List)
Biology
ISBN:9781305389892
Author:Peter J. Russell, Paul E. Hertz, Beverly McMillan
Publisher:Cengage Learning

Surgical Tech For Surgical Tech Pos Care
Health & Nutrition
ISBN:9781337648868
Author:Association
Publisher:Cengage

Essentials of Pharmacology for Health Professions
Nursing
ISBN:9781305441620
Author:WOODROW
Publisher:Cengage
genetic recombination strategies of bacteria CONJUGATION, TRANSDUCTION AND TRANSFORMATION; Author: Scientist Cindy;https://www.youtube.com/watch?v=_Va8FZJEl9A;License: Standard youtube license