
Concept explainers
A study was conducted in an attempt to determine which functional regions of a particular conjugative transfer gene (tral) are involved in the transfer of plasmid R27 in Salmonella enterica. The R27 plasmid is of significant clinical interest because it is capable of encoding multiple-antibiotic resistance to typhoid fever. To identify functional regions responsible for conjugal transfer, an analysis by Lawley et al. [(2002). J. Bacteriol. 184:2173-2180] was conducted in which particular regions of the tral gene were mutated and tested for their impact on conjugation. Shown here is a map of the regions tested and believed to be involved in conjugative transfer of the plasmid. Similar coloring indicates related function. Numbers correspond to each functional region subjected to mutation analysis.
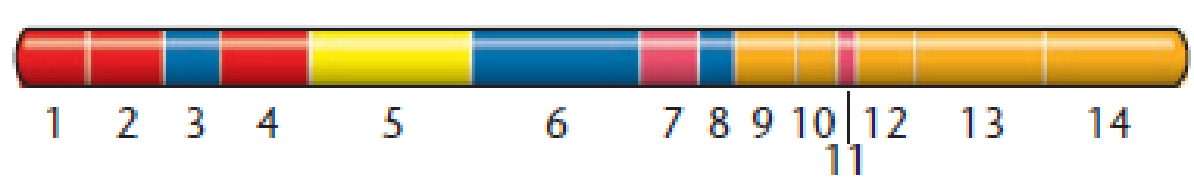
Accompanying the map is a table showing the effects of these mutations on R27 conjugation.
Effects of Mutations in Functional Regions of Transfer Region 1 (tral) on R27 Conjugation
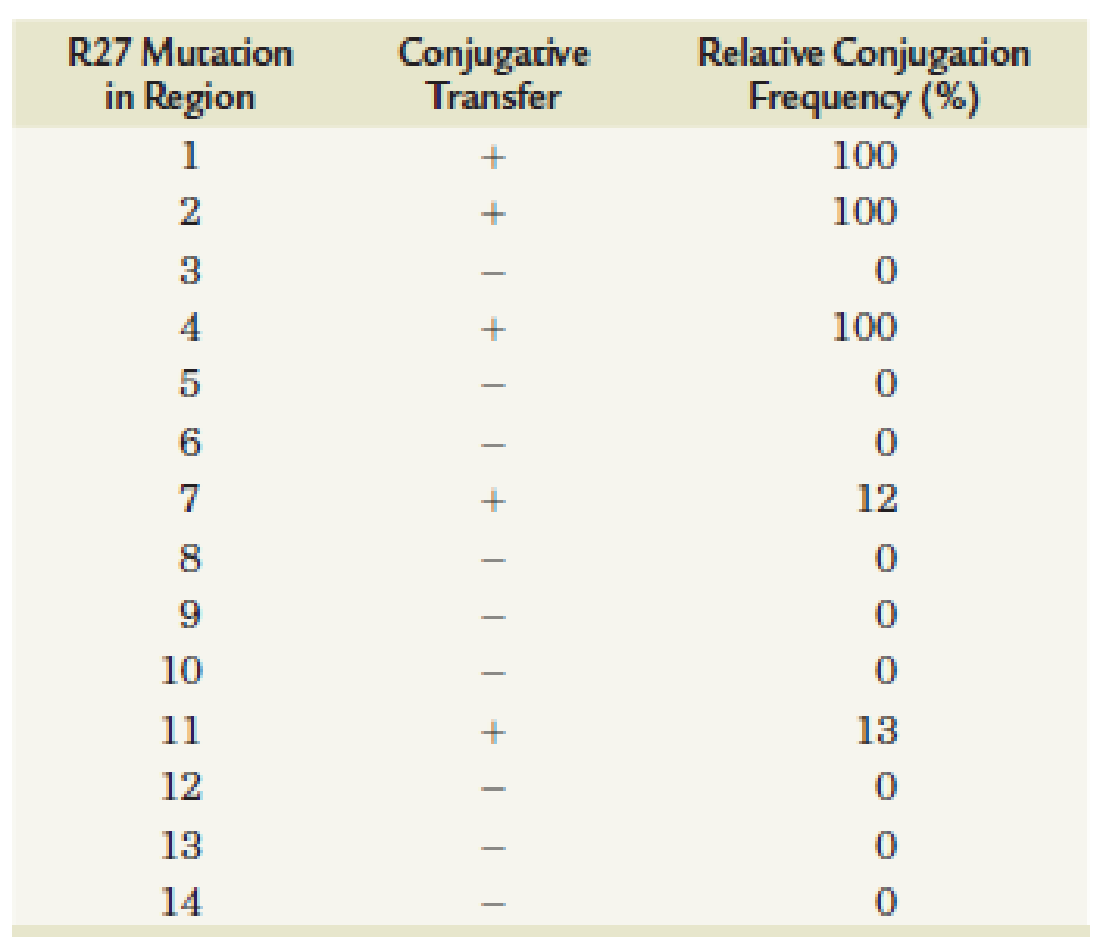
- (a) Given the data, do all functional regions appear to influence conjugative transfer?
- (b) Which regions appear to have the most impact on conjugation?
- (c) Which regions appear to have a limited impact on conjugation?
- (d) What general conclusions might one draw from these data?
Want to see the full answer?
Check out a sample textbook solution
Chapter 6 Solutions
Concepts of Genetics (12th Edition)
- 18. Watch this short youtube video about SARS CoV-2 replication. SARS-CoV-2 Life Cycle (Summer 2020) - YouTube.19. What is the name of the receptor that SARS CoV-2 uses to enter cells? Which human cells express this receptor? 20. Name a few of the proteins that the SARS CoV-2 mRNA codes for. 21. What is the role of the golgi apparatus related to SARS CoV-2arrow_forwardState the five functions of Globular Proteins, and give an example of a protein for each function.arrow_forwardDiagram of check cell under low power and high powerarrow_forward
- a couple in which the father has the a blood type and the mother has the o blood type produce an offspring with the o blood type, how does this happen? how could two functionally O parents produce an offspring that has the a blood type?arrow_forwardWhat is the opening indicated by the pointer? (leaf x.s.) stomate guard cell lenticel intercellular space none of thesearrow_forwardIdentify the indicated tissue? (stem x.s.) parenchyma collenchyma sclerenchyma ○ xylem ○ phloem none of thesearrow_forward
- Where did this structure originate from? (Salix branch root) epidermis cortex endodermis pericycle vascular cylinderarrow_forwardIdentify the indicated tissue. (Tilia stem x.s.) parenchyma collenchyma sclerenchyma xylem phloem none of thesearrow_forwardIdentify the indicated structure. (Cucurbita stem l.s.) pit lenticel stomate tendril none of thesearrow_forward
- Identify the specific cell? (Zebrina leaf peel) vessel element sieve element companion cell tracheid guard cell subsidiary cell none of thesearrow_forwardWhat type of cells flank the opening on either side? (leaf x.s.) vessel elements sieve elements companion cells tracheids guard cells none of thesearrow_forwardWhat specific cell is indicated. (Cucurbita stem I.s.) vessel element sieve element O companion cell tracheid guard cell none of thesearrow_forward
 Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Concepts of BiologyBiologyISBN:9781938168116Author:Samantha Fowler, Rebecca Roush, James WisePublisher:OpenStax College
Concepts of BiologyBiologyISBN:9781938168116Author:Samantha Fowler, Rebecca Roush, James WisePublisher:OpenStax College





