
Concept explainers
An Hfr strain is used to map three genes in an interrupted mating experiment. The cross is Hfr/a+b+c+rif × F−>a−b−c−rifr. (No map order is implied in the listing of the alleles; rifr is resistance to the antibiotic rifampicin.) The a+ gene is required for the biosynthesis of nutrient A, the b+ gene for nutrient B, and c+ for nutrient C. The minus alleles are auxotrophs for these nutrients. The cross is initiated at time = 0, and at various times, the mating mixture is plated on three types of medium. Each plate contains minimal medium (MM) plus rifampicin plus specific supplements that are indicated in the following table. (The results for each time interval are shown as the number of colonies growing on each plate.)
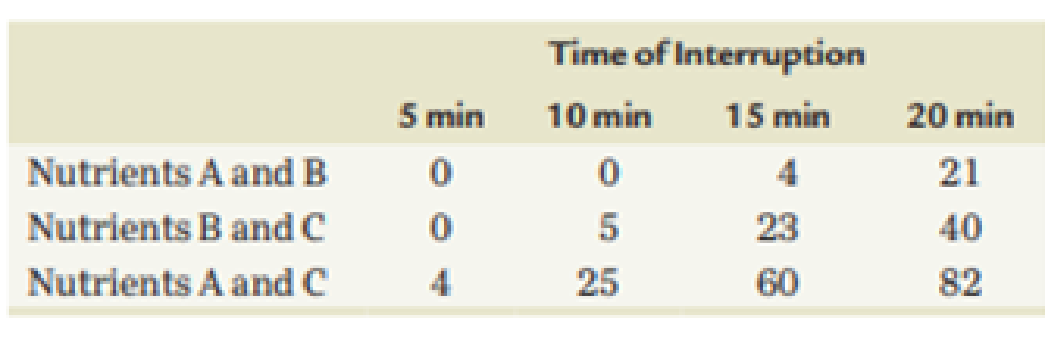
- (a) What is the purpose of rifampicin in the experiment?
- (b) Based on these data, determine the approximate location on the chromosome of the a, b, and c genes relative to one another and to the F factor.
- (c) Can the location of the rif gene be determined in this experiment? If not, design an experiment to determine the location of rif relative to the F factor and to gene b.
Want to see the full answer?
Check out a sample textbook solution
Chapter 6 Solutions
Concepts of Genetics (12th Edition)
- the direct output from molecular replacement is a coordinate file showing the orientation of the unknown target protein in the unit cell. true or false?arrow_forwardDid your intake of vitamin C meet or come very close to the recommended amount? yes noarrow_forwardWhich of the following statements about hydration is true? Absence of thirst is a reliable indication that an individual is adequately hydrated. All of these statements are true. Although a popular way to monitor hydration status, weighing yourself before and after intensive physical activity is not a reliable method to monitor hydration. Urine that is the color of apple juice indicates dehydration. I don't know yetarrow_forward
- Three of the many recessive mutations in Drosophila melanogaster that affect body color, wing shape, or bristle morphology are black (b) body versus grey in wild type, dumpy (dp), obliquely truncated wings versus long wings in the male, and hooked (hk) bristles versus not hooked in the wild type. From a cross of a dumpy female with a black and hooked male, all of the F1 were wild type for all three of the characters. The testcross of an F1 female with a dumpy, black, hooked male gave the following results: Trait Number of individuals Wild type 169 Black 19 Black, hooked 301 Dumpy, hooked 21 Hooked, dumpy, black 172 Dumpy, black 6 Dumpy 305 Hooked 8 Determine the order of the genes and the mapping distance between genes. Determine the coefficient of confidence for the portion of the chromosome involved in the cross. How much interference takes place in the cross?arrow_forwardWhat happens to a microbes membrane at colder temperature?arrow_forwardGenes at loci f, m, and w are linked, but their order is unknown. The F1 heterozygotes from a cross of FFMMWW x ffmmww are test crossed. The most frequent phenotypes in the test cross progeny will be FMW and fmw regardless of what the gene order turns out to be. What classes of testcross progeny (phenotypes) would be least frequent if locus m is in the middle? What classes would be least frequent if locus f is in the middle? What classes would be least frequent if locus w is in the middle?arrow_forward
- 1. In the following illustration of a phospholipid... (Chemistry Primer and Video 2-2, 2-3 and 2-5) a. Label which chains contain saturated fatty acids and non-saturated fatty acids. b. Label all the areas where the following bonds could form with other molecules which are not shown. i. Hydrogen bonds ii. Ionic Bonds iii. Hydrophobic Interactions 12-6 HICIH HICIH HICHH HICHH HICIH OHHHHHHHHHHHHHHHHH C-C-C-C-C-c-c-c-c-c-c-c-c-c-c-c-C-C-H HH H H H H H H H H H H H H H H H H H HO H-C-O H-C-O- O O-P-O-C-H H T HICIH HICIH HICIH HICIH HHHHHHH HICIH HICIH HICIH 0=C HIC -C-C-C-C-C-C-C-C-CC-C-C-C-C-C-C-C-C-H HHHHHHHHH IIIIIIII HHHHHHHH (e-osbiv)arrow_forwardAnswer this as a dental assistant studentarrow_forwardbuatkan judul skripsi tentang parasitologi yang sedang trendinharrow_forward
- Dental assistantarrow_forwardO Macmillan Learning Glu-His-Trp-Ser-Gly-Leu-Arg-Pro-Gly The pKa values for the peptide's side chains, terminal amino groups, and carboxyl groups are provided in the table. Amino acid Amino pKa Carboxyl pKa Side-chain pKa glutamate 9.60 2.34 histidine 9.17 1.82 4.25 6.00 tryptophan 9.39 2.38 serine 9.15 2.21 glycine 9.60 2.34 leucine 9.60 2.36 arginine 9.04 2.17 12.48 proline 10.96 1.99 Calculate the net charge of the molecule at pH 3. net charge at pH 3: Calculate the net charge of the molecule at pH 8. net charge at pH 8: Calculate the net charge of the molecule at pH 11. net charge at pH 11: Estimate the isoelectric point (pl) for this peptide. pl:arrow_forwardBiology Questionarrow_forward
 Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning- Case Studies In Health Information ManagementBiologyISBN:9781337676908Author:SCHNERINGPublisher:Cengage
 Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning





