
Concept explainers
Imagine you have found a small quantity of DNA. Fill in the following diagram, which outlines a series of DNA technology experiments you could perform to study this DNA.
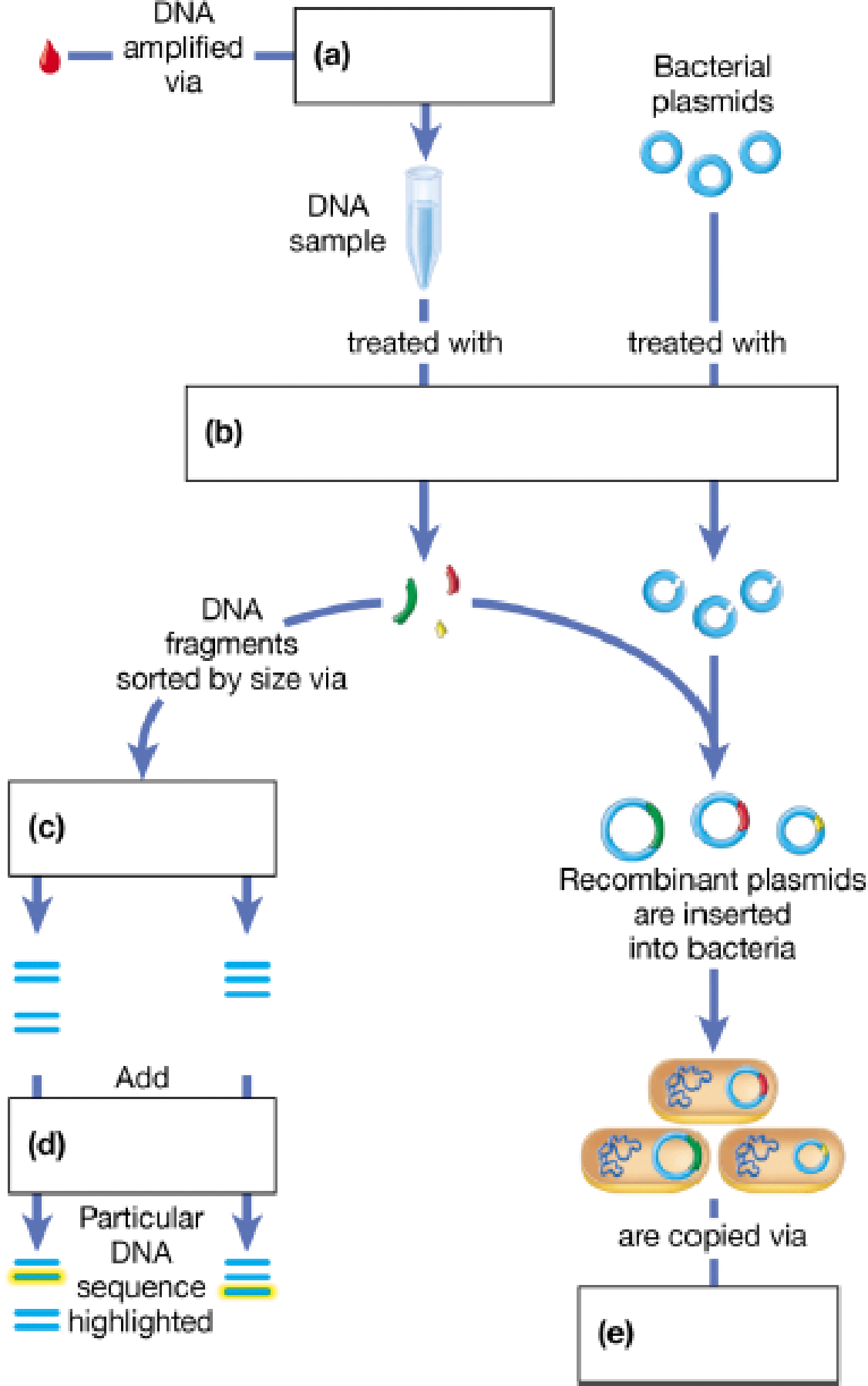
To complete: The given map showing the steps involved in recombinant DNA (rDNA) technology.
Introduction: Recombinant DNA technology is a technique used to produce novel genetic combinations by combining the genes from two different organisms and introduces them inside a host in order to proliferate and grow.
Answer to Problem 1CC
Pictorial representation: Fig. 1 shows the completed map of the steps involved in recombinant DNA technology.
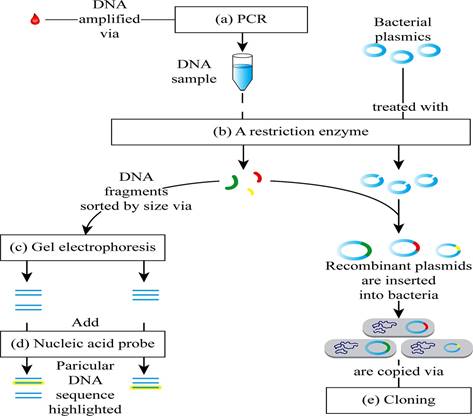
Fig. 1 The steps involved in recombinant DNA (rDNA) technology.
Explanation of Solution
(a)
Correct answer: PCR
Explanation: The polymerase chain reaction (PCR) is used to get multiple copies of DNA within a short period. In PCR, for synthesizing multiple copies of a specific DNA fragment, DNA nucleotide primers are used. Hence, the correct answer is PCR.
(b)
Correct answer: A restriction enzyme
Explanation: Restriction enzymes are a group of enzymes used to cut the DNA by binding to certain specific DNA sequence sites called as restriction sites. It forms single-stranded ends called as sticky ends. The sticky ends help the DNA to bind with the other DNA fragment that has the same single-stranded ends by complementary base pairing. Hence, the correct answer is a restriction enzyme.
(c)
Correct answer: Gel electrophoresis
Explanation: Electrophoresis is used to separate the molecules based on their size and charge in the presence of an electric field. Hence, the correct answer is Gel electrophoresis.
(d)
Correct answer: Nucleic acid probe
Explanation: Nucleic acid probe is used for the analysis of the probe, which can specifically bind to the complementary DNA strand. Hence, the correct answer is nucleic acid probe.
(e)
Correct answer: Cloning
Explanation: The process of production of identical copies of the parent cell is called as cloning. Cloning vectors are used in the rDNA technology for the propagation of the desired gene. Hence, the correct answer is cloning.
Want to see more full solutions like this?
Chapter 12 Solutions
Campbell Biology: Concepts & Connections (8th Edition)
- Please answer the following chart so I can understand how to do it.arrow_forwardDigoxin: Intravenous Bolus - Two Compartment Model Drug Digoxin Route: IV Bolus Dose: 0.750 mg Plasma Concentration Time Profile Beta Alpha Time (hrs) Conc (ng/ml) LN (ng/ml) LN (ng/ml) LN 0.00 #NUM! #NUM! #NUM! 0.10 12.290 2.509 #NUM! #NUMI 0.60 6.975 1.942 #NUM! #NUMI 1.00 4.649 1.537 #NUM! #NUMI 2.00 2.201 0.789 #NUM! #NUM! 3.00 1.536 0.429 #NUM! #NUM! 4.00 1.342 0.294 #NUM! #NUM! 5.00 1.273 0.241 #NUM! #NUMI 6.00 1.238 0.213 #NUM! #NUM! 7.00 1.212 0.192 #NUM! #NUM! 8.00 1.188 0.172 #NUMI #NUM! 9.00 1.165 0.153 #NUM! #NUMI 10.00 1.143 0.134 #NUMI #NUM! 11.00 1.122 0.115 #NUM! 12.00 1.101 0.096 #NUMI 13.00 1.080 0.077 #NUMI 16.00 1.020 0.020 #NUMI 24.00 0.876 -0.132 #NUMI Pharmacokinetic Parameters Parameter Value Alpha B Beta Units ng/ml hr-1 ng/ml hr-1 CO ng/ml H.C AUC ng x hr/ml Vc Vbeta Vss C L/hr TK (alpha) hr TX (beta) days 5+ F3 F4 F5 0+ F6 F7 % 6 95 14 #3 29 & t F8 F9 FW EWarrow_forwardLinuron, a derivative of urea, is used as an herbicide. Linuron serum levels were measured in 4Kg rabbits following a bolus IV injection of 10mg/kg. Time (minutes) Serum Linuron Levels (ug/ml) following IV dose 10 15.48 20 8.60 30 5.90 45 3.78 60 2.42 90 1.49 120 0.93 180 0.60 240 0.41 300 0.29 360 0.22 Analyze this data and perform the necessary calculations to determine the following pharmacokinetic parameters from the IV data: (5 points per parameter, 24 parameters/variables ■ 120 points possible). You do NOT need to submit graphs or data tables. Give the terminal regression line equation and R or R² value: Give the x axis (name and units, if any) of the terminal line: Give the y axis (name and units, if any) of the terminal line: Give the residual regression line equation and R or R² value: Give the x axis (name and units, if any) of the residual line: Give the y axis (name and units, if any) of the residual line:arrow_forward
- 3. In the tomato, red fruit (O+) is dominant over orange fruit (0), and yellow flowers (W+) are dominant over white flowers (w). A cross was made between true-breeding plants with red fruit and yellow flowers, and plants with orange fruit and white flowers. The F₁ plants were then crossed to plants with orange fruit and white flowers, which produced the following results: a. b. 333 red fruit, yellow flowers 64 red fruit, white flowers 58 orange fruit, yellow flowers 350 orange fruit, white flowers Conduct a chi-square analysis to demonstrate that these two genes DO NOT assort independently. Make sure to interpret the P value obtained from your chi-square test. Calculate and provide the map distance (in map units) between the two genes.arrow_forwardName: Date: Investigation: Is a dog more closely related to a coyote or a wolf? Gray Wolf Species Name: Canis lupus Color: Light gray to black Size: 80-120 pounds, 2.5 feet tall Appearance: broad snout, round ears, long tail Coyote Species Name: Canis latrans Color: Light gray to brown Size: 20-50 pounds, 1.5 feet tall Appearance: narrow snout, pointed ears, long tail Dog, Alaskan Malamute Species Name: Canis lupus familiaris Color: Gray and white or brown and white Size: 70-80 pounds, 2 feet tall Appearance: broad snout, round ears, long tail 1. Examine the images and descriptions above. Underline any similarities between the dog and the wolf. Place a star next to any coyote traits that are similar to the dog. 2. Based on appearance alone, which do you think is the most closely related to a dog? www.biologycorner.comarrow_forwardyu yeuwyuyuierydtgcygucygzycghjcygyugfyudguygcywgduycgyudgs ygarrow_forward
- According to a recent study, 1 out of 50,000 people will be diagnosed with cystic fibrosis. Cystic fibrosis can be caused by a mutant form of the CFTR gene (dominant gene symbol is F and mutant is f). A. Using the rate of incidence above, what is the frequency of carriers of the cystic fibrosis allele for CFTR in the US? (3 pts) B. In a clinical study, 400 people from the population mentioned in (A.) were genotyped for BRCA1 Listed below are the results. Are these results in Hardy- Weinberg equilibrium? Use Chi Square to show whether or not they are. (3 pts) # of women BRCA1 genotype BB 390 Bb bb 12pt v 10 0 V Paragraph B IUA BIUA > V T² v <arrow_forwardCase Study—Ella Ella has a family history of diabetes. She wants to follow a healthful eating pattern that can lower her risk for developing this condition. Her dietitian recommends a goal of 450 to 600 kcal per meal and advises Ella to follow the Acceptable Macronutrient Distribution Range (AMDR) for carbohydrates and the Dietary Guidelines for Americans 2015-2020, which recommend limiting added sugar. She also recommends that Ella choose whole grains rather than processed grains. Ella decides to pack a lunch to take to work every day. This morning she’s making a sandwich for her lunch. Categories of Sandwich Options (Top of the screen) Breads Spreads Cheeses Vegetables Proteins Specific food items to select White Bread 6-inches Honey Mustard Provolone LettuceTomatoBell Peppers Turkey Part A - Reading Nutrition Facts Panels for Total Kilocalories How many total kilocalories are in Ella’s sandwich? _______ kcal ? Part B - Reading Nutrition Facts Panels for…arrow_forwardCase Study—Ella Ella has a family history of diabetes. She wants to follow a healthful eating pattern that can lower her risk for developing this condition. Her dietitian recommends a goal of 450 to 600 kcal per meal and advises Ella to follow the Acceptable Macronutrient Distribution Range (AMDR) for carbohydrates and the Dietary Guidelines for Americans 2015-2020, which recommend limiting added sugar. She also recommends that Ella choose whole grains rather than processed grains. Ella decides to pack a lunch to take to work every day. This morning she’s making a sandwich for her lunch. Categories of Sandwich Options (Top of the screen) Breads Spreads Cheeses Vegetables Proteins Specific food items to select White Bread 6-inches Honey Mustard Provolone LettuceTomatoBell Peppers Turkey Part A - Reading Nutrition Facts Panels for Total Kilocalories How many total kilocalories are in Ella’s sandwich exactl ______kcal ? Part B - Reading Nutrition Facts Panels for…arrow_forward
- In humans, red-green color blindness is recessive and X-linked, whereas albinism is recessive and autosomal. What types of children can be produced as the result of marriage between two homozygous parents, a normal-vision albino woman and a color-blind, normal male?arrow_forwardIn Drosophila, an X linked recessive mutation, scalloped (sd), causes irregular wing margins. Diagram the F1 and F2 results if a (a) scalloped female is crossed with a normal male; (b) a scalloped male is crossed with a normal female (assume the female is homozygous). Compare these results to what you would find if the trait was not sex linked.arrow_forwardCase Study—Ella Ella has a family history of diabetes. She wants to follow a healthful eating pattern that can lower her risk for developing this condition. Her dietitian recommends a goal of 450 to 600 kcal per meal and advises Ella to follow the Acceptable Macronutrient Distribution Range (AMDR) for carbohydrates and the Dietary Guidelines for Americans 2015-2020, which recommend limiting added sugar. She also recommends that Ella choose whole grains rather than processed grains. Ella decides to pack a lunch to take to work every day. This morning she’s making a sandwich for her lunch. Categories of Sandwich Options (Top of the screen) Breads Spreads Cheeses Vegetables Proteins Specific food items to select White Bread 6-inches Honey Mustard Provolone LettuceTomatoBell Peppers Turkey Part A - Reading Nutrition Facts Panels for Total Kilocalories How many total kilocalories are in Ella’s sandwich exactl ______kcal ? Part B - Reading Nutrition Facts Panels for…arrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning
Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning Concepts of BiologyBiologyISBN:9781938168116Author:Samantha Fowler, Rebecca Roush, James WisePublisher:OpenStax College
Concepts of BiologyBiologyISBN:9781938168116Author:Samantha Fowler, Rebecca Roush, James WisePublisher:OpenStax College Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax





