
Concept explainers
To analyze:
The six nucleotides prior to the start codon and the first
Introduction:
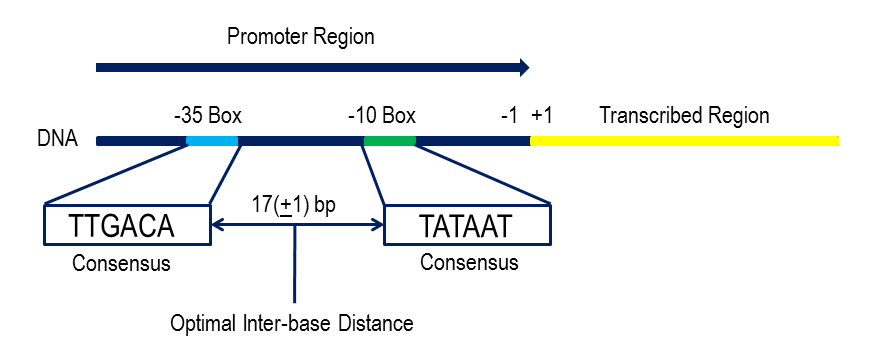
Consensus sequences are the short stretch of DNA sequences that are commonly located nucleotides found at a specific location of DNA and RNA or amino acids. These sequences are used for inter or intramolecular interactions. These sequences are similar to structure and functions in all organisms. The consensus sequence is a calculated order that states the most frequent nucleotide at that position in the alignment.
Want to see the full answer?
Check out a sample textbook solution
Chapter 9 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- Ch.21 What causes patients infected with the yellow fever virus to turn yellow (jaundice)? A. low blood pressure and anemia B. excess leukocytes C. alteration of skin pigments D. liver damage in final stage of disease — What is the advantage for malarial parasites to grow and replicate in red blood cells? A. able to spread quickly B. able to avoid immune detection C. low oxygen environment for growth D. cooler area of the body for growth — Which microbe does not live part of its lifecycle outside humans? A. Toxoplasma gondii B. Cytomegalovirus C. Francisella tularensis D. Plasmodium falciparum — explain your answer thoroughlyarrow_forwardCh.22 Streptococcus pneumoniae has a capsule to protect it from killing by alveolar macrophages, which kill bacteria by… A. cytokines B. antibodies C. complement D. phagocytosis — What fact about the influenza virus allows the dramatic antigenic shift that generates novel strains? A. very large size B. enveloped C. segmented genome D. over 100 genes — explain your answer thoroughlyarrow_forwardWhat is this?arrow_forward
- Molecular Biology A-C components of the question are corresponding to attached image labeled 1. D component of the question is corresponding to attached image labeled 2. For a eukaryotic mRNA, the sequences is as follows where AUGrepresents the start codon, the yellow is the Kozak sequence and (XXX) just represents any codonfor an amino acid (no stop codons here). G-cap and polyA tail are not shown A. How long is the peptide produced?B. What is the function (a sentence) of the UAA highlighted in blue?C. If the sequence highlighted in blue were changed from UAA to UAG, how would that affecttranslation? D. (1) The sequence highlighted in yellow above is moved to a new position indicated below. Howwould that affect translation? (2) How long would be the protein produced from this new mRNA? Thank youarrow_forwardMolecular Biology Question Explain why the cell doesn’t need 61 tRNAs (one for each codon). Please help. Thank youarrow_forwardMolecular Biology You discover a disease causing mutation (indicated by the arrow) that alters splicing of its mRNA. This mutation (a base substitution in the splicing sequence) eliminates a 3’ splice site resulting in the inclusion of the second intron (I2) in the final mRNA. We are going to pretend that this intron is short having only 15 nucleotides (most introns are much longer so this is just to make things simple) with the following sequence shown below in bold. The ( ) indicate the reading frames in the exons; the included intron 2 sequences are in bold. A. Would you expected this change to be harmful? ExplainB. If you were to do gene therapy to fix this problem, briefly explain what type of gene therapy youwould use to correct this. Please help. Thank youarrow_forward
- Molecular Biology Question Please help. Thank you Explain what is meant by the term “defective virus.” Explain how a defective virus is able to replicate.arrow_forwardMolecular Biology Explain why changing the codon GGG to GGA should not be harmful. Please help . Thank youarrow_forwardStage Percent Time in Hours Interphase .60 14.4 Prophase .20 4.8 Metaphase .10 2.4 Anaphase .06 1.44 Telophase .03 .72 Cytukinesis .01 .24 Can you summarize the results in the chart and explain which phases are faster and why the slower ones are slow?arrow_forward
- Can you circle a cell in the different stages of mitosis? 1.prophase 2.metaphase 3.anaphase 4.telophase 5.cytokinesisarrow_forwardWhich microbe does not live part of its lifecycle outside humans? A. Toxoplasma gondii B. Cytomegalovirus C. Francisella tularensis D. Plasmodium falciparum explain your answer thoroughly.arrow_forwardSelect all of the following that the ablation (knockout) or ectopoic expression (gain of function) of Hox can contribute to. Another set of wings in the fruit fly, duplication of fingernails, ectopic ears in mice, excess feathers in duck/quail chimeras, and homeosis of segment 2 to jaw in Hox2a mutantsarrow_forward
 Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage LearningEssentials Health Info Management Principles/Prac...Health & NutritionISBN:9780357191651Author:BowiePublisher:Cengage
Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage LearningEssentials Health Info Management Principles/Prac...Health & NutritionISBN:9780357191651Author:BowiePublisher:Cengage





