
Concept explainers
Yeast are single-celled eukaryotic organisms that grow in culture as either haploids or diploids. Diploid yeast are generated when two haploid strains fuse together. Seven haploid strains of yeast exhibit similar growth habit: At
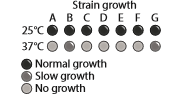
Describe the nature of the mutation affecting each of these mutant yeast strains. Explain why strains B and G display different growth habit at
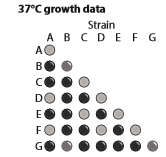

How many different genes are mutated among these seven yeast strains? Identify the strains that represent each gene mutation.
Want to see the full answer?
Check out a sample textbook solution
Chapter 4 Solutions
Pearson eText Genetic Analysis: An Integrated Approach -- Instant Access (Pearson+)
- There are five substitution mutations in the dark-colored mutant Mc1r gene. Compare the DNA sequence of the light-colored wild-type Mc1r gene with the DNA sequence of the dark-colored mutant Mc1r gene. Indicate the locations of the five mutations by changing the font color to YELLOW for the five single DNA nucleotides that are mutated in the mutant Mc1r gene table. Using the information in the introduction, determine whether each of these mutations is a silent, missense, or nonsense mutation. Using the mutant Mc1r gene data, fill in the columns (including DNA, mRNA, and amino acid) in gene table 2 that contain a silent mutation with BLUE. Likewise, fill in the columns that contain a missense mutation with RED. Shade any columns that contain nonsense mutations with GREEN. Then Of the five mutations you identified in the mutant Mc1r gene, how many are: substitutions insertions deletions (Enter a number on each line.) 2. Of the five mutations…arrow_forwardA series of Hfr strains that have genotype m+ n+ o+ p+ q+ r+ are mixed with an F− strain that has genotype m− n− o− p− q− r−. Conjugation is interrupted at regular intervals and the order of the appearance of genes from the Hfr strain is determined in the recipient cells. The order of gene transfer for each Hfr strain is What is the order of genes on the circular bacterial chromosome? For each Hfr strain, give the location of the F factor in the chromosome and its polarity. Hfr5 m+ q+ p+ n+ r+ o+ Hfr4 n+ r+ o+ m+ q+ p+ Hfr1 o+ m+ q+ p+ n+ r+ Hfr9 q+ m+ o+ r+ n+ p+arrow_forwardYou have isolated 8 mutants in yeast that fail to grow on minimal media plates but do grow when they are supplemented with Arginine. You know that Arginine is synthesized in a biochemical pathway within wild-type yeast, but you do not know how many gene products it takes for the pathway. You have all of the lines as both a and a cells and mate each strain to each other in pairwise crosses and plate them on minimal media to see if they grow. You obtain the following results with (+) representing growth, and (-) indicating no growth: a 1 5 1 a 4 5 6 7 8 How many genes are represented? O 1 3 7 O Cannot tell from the data a + + + + + • + + i 0 +, + + + • + + 7 + + + + + , . + + + + + m + + + + + + + 2 + + + + + i + -I + + . . + + +arrow_forward
- You have a strain of Neurospora that is unable to synthesize histidine and thus requires H in the media in order to grow. You have isolated one revertant colony. Predict the expected proportion of the progeny that would be h+ if you cross the colony with the original mutant colony and the reversion occurred by each of the following mechanisms: Precise change of the mutated base back to its original base. A suppressor gene is mutated on a different chromosome A suppressor gene is mutated on the same chromosome but 10mu distant from the mutated gene. The mutant colony is crossed to a wild-type Neurospora colony and the following data are collected. 95% of all asci scored are h+ but 5% are h-. Which mechanism in part a is consistent with these data? Explain why and what has happened on a molecular level.arrow_forwardAssume that a series of compounds has been discovered in Neurospora. Compounds A–F appear to be intermediates in a biochemical pathway. Conversion of one intermediate to the next is controlled by enzymes that are encoded by genes. Several mutations in these genes have been identified and Neurospora strains 1–4 each contain a single mutation. Strains 1–4 are grown on minimal media supplemented with one of the compounds A–F. The ability of each strain to grow when supplemented with different compounds is shown in the table (+ = growth; o = no growth). Which biochemical pathway fits the data presented? Media Supplement Strain A B C D E F 1 o o o + + + 2 o o o o + + 3 o o o o + o 4 o o + + + + A) A → B → C → D → E → F B) A → B → C → F → D → E C) F → B → C → D → A → E D) A → B → C → D → F → E E) A → B → F → E → C → Darrow_forwardA yeast geneticist irradiates haploid cells of a strain thatis an adenine-requiring auxotrophic mutant, caused bymutation of the gene ade1. Millions of the irradiatedcells are plated on minimal medium, and a small number of cells divide and produce prototrophic colonies.These colonies are crossed individually with a wildtype strain. Two types of results are obtained:(1) prototroph × wild type : progeny all prototrophic(2) prototroph × wild type : progeny 75% prototrophic,25% adenine-requiring auxotrophsa. Explain the difference between these two types ofresults.b. Write the genotypes of the prototrophs in each case.c. What progeny phenotypes and ratios do you predictfrom crossing a prototroph of type 2 by the original ade1auxotroph?arrow_forward
- There are five substitution mutations in the dark-colored mutant Mc1r gene. Compare the DNA sequence of the light-colored wild-type Mc1r gene with the DNA sequence of the dark-colored mutant Mc1r gene. Indicate the locations of the five mutations by changing the font color to YELLOW for the five single DNA nucleotidesthat are mutated in the mutant Mc1r gene table. Using the information in the introduction, determine whether each of these mutations is a silent, missense, or nonsense mutation. Using the mutant Mc1r gene data, fill in the columns (including DNA, mRNA, and amino acid) in gene table 2 that contain a silent mutation with BLUE. Likewise, fill in the columns that contain a missense mutation with RED. Shade any columns that contain nonsense mutations with GREEN.arrow_forwardBrenner’s m mutant phages (m1–m6) described inFig. 8.8 were suppressed when grown in suppressor(su−) mutant bacteria; they produced full-length Mproteins that functioned like wild-type M protein.a. What gene do you think was mutant in the su−bacteria?b. When the m− phages were propagated in the su−bacterial strain, not all of the proteins made by themutant m alleles were identical to wild-type Mprotein. How did some of them differ?arrow_forwardA shuttle vector is a vector (usually a plasmid) constructed so that it can propagate in two different host species. One of the most common types of shuttle vectors is the yeast shuttle vector. Examples of such vectors derived from yeast are Yeast Episomal Plasmid (YEp), Yeast Integrating Plasmid (YIp) and Yeast Replicating Plasmid (YRp). Among these three vectors, YIp has the lowest transformation frequency and copy number per cell. Explain why Ylp is still popularly used despite its limitations.arrow_forward
- The figure below shows a partial chromosome map of an E. coli Hfr strain. Each mark = 10 minutes between conjugation transfer time. If transfer of genes begins at “*” relative to the origin of transfer, what is one of the predicted results from this map? It would take less than 30 minutes to transfer all of the genes that are shown. gal and azi will rarely be transferred together. gal and ton will rarely be transferred together. Ten minutes after transfer of ton, lac will be transferred. This strain will produce very few gal recombinants.arrow_forwardThe presence (+) or absence (−) of six sequences in each of five bacterial artificial chromosome (BAC) clones (A–E) is indicated in the following table. Using these markers, put the BAC clones in their correct order and indicate the locations of the numbered sequences within them. Sequences BAC clone 1 2 3 4 5 6 A + − − − + − B − − − + − + C − + + − − − D − − + − + − E + − − + − −arrow_forwardMost yeast grow suspended in culture, but some yeast grow as a film on the top of a liquid culture. These are called flor yeast. The following Southern blot is looking at the FLO11 gene that is thought to be involved in whether yeast are flor or not. DNA was isolated from four different yeast strains. Strains 1 and 3 are normal while strains 2 and 4 are flor (Fidalgo, 2006). Based on the gene map and Southern results below, what aspect of FLO11 determines whether a strain will be flor or not?arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





