
Concept explainers
To further analyze the CRABS CLAW gene (see Problems

What restriction digest would help resolve any ambiguity in the map?
You have isolated a genomic clone with an EcoRI fragment of
Does this tell you anything about where the CRABS CLAW gene is located within the 11-kb genomic clone?
Restriction enzyme sites within a cDNA clone are often also found in the genomic sequence. Can you think of a reason why occasionally this is not the case? What about the converse: Are restriction enzyme sites in a genomic clone always in a cDNA clone of the same gene?
You have identified a
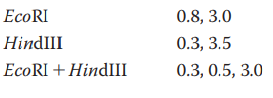
Want to see the full answer?
Check out a sample textbook solution
Chapter 17 Solutions
GENETIC ANALYSIS: AN INTEG. APP. W/MAS
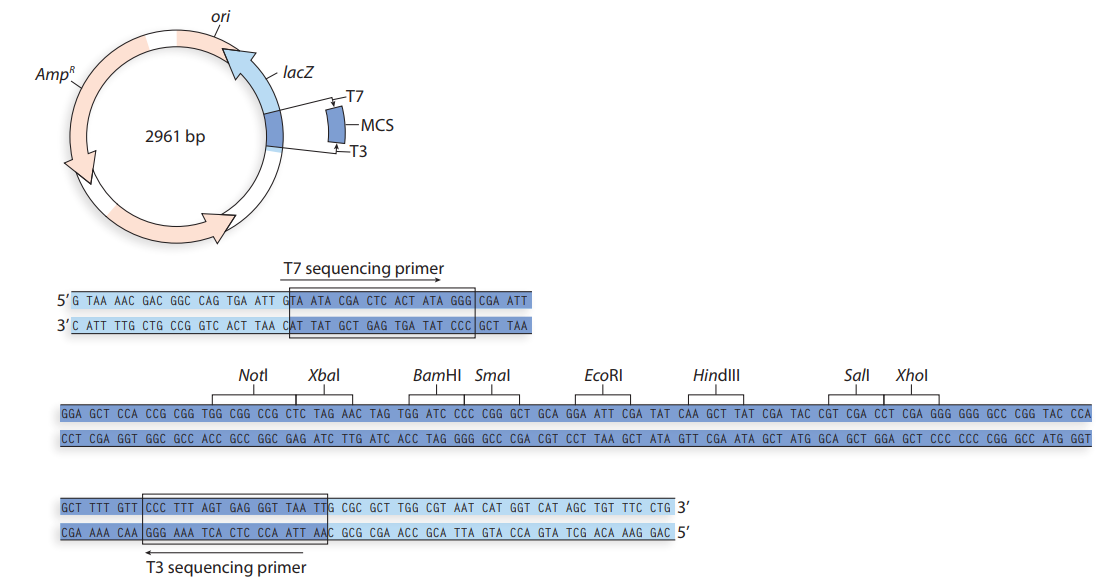
- Knowing that you are using HindIII and EcoRI to cut your plasmids, and that those two enzymes cut within the MCS, use the map of pUC19 provided below to compute: What will be the sizes of the 2 restriction fragments if NO insert is present in pUC19? What will be the sizes of the 2 restriction fragments if the approximately 317 bp RT-PCR product (insert from WT satC dimer) was ligated successfully into the SmaI site? What will be the sizes of the 2 restriction fragments if TWO approximately 317 bp RT-PCR products (2 ligated inserts from WT satC dimer) were ligated into the SmaI site?arrow_forwardA group of overlapping clones, designated A through F, is isolated from one region of a chromosome. Each of the clones is separately cleaved by a restriction enzyme, and the pieces are resolved by agarose gel lectrophoresis,with the results shown below. There are nine different restriction fragments in this chromosomal region, with a subset appearing in each clone. Using this information, deduce the order of the restriction fragments in the chromosome.arrow_forwardDecide on two restriction sites that you can use to clone this into pL4440’s MCS. Identify their sequence. Tip: The plasmid map is in Figure 3, details of restriction site sequences can be found at https://enzymefinder.neb.com/#!#nebheaderarrow_forward
- 1) You wish to make a restriction map of a 17.0 kb linear fragment. You digest the fragment with Sbf1, Pst1, and a mixture of Sbf1 and Pst1. From these results, you obtain these fragments following agarose gel electrophoresis of the three samples: Sbf1: 7kb and 10kb Pstl: 3kb, 6kb and 8kb Sbf1 and Pstl: 1kb, 2kb, 6kb and 8kb Question: In which of the Pstl fragments is the Sbfl site located? 8kb O 6kb 3kb O None of the abovearrow_forwardIn making recombinant DNA molecules that combine restriction fragments from different organisms, researchers usually prefer restriction enzymes like BamHI or HindIII that generate fragments with “sticky ends” (ends with overhangs) rather than enzymes like HpaI or SmaI (Table 12.1) that generate fragments with “blunt ends” (ends without overhangs). Can you think of a reason for this preference?arrow_forwardAfter characterizing the DNA composition of various cats, you identify a protein-coding gene in tigers called stripes and wish to study the structure of the protein product STRIPES. This requires that you purify recombinant ridges from E. coli. First, the stripes gene must be amplified by PCR and then inserted into an appropriate plasmid for bacterial expression. Such a plasmid is diagrammed below. ori CAP Binding Site من Promoter MCS The restriction sites for Aatll and Kpnl are: Aatll 5'-GACGTC-3' Kpnl = 5'-GGTACC-3' Laco (Operator) -Kpnl Aatll The coding strand for the stripes gene is shown below, with start and stop codons in bold. 5'-ATGCAACAGTAGCTGAAGCCCAGTGACACCATCGAAAATGTGAAGGCCAAGATGAGGCTCATCTTTGCAGGCAAGCAGCTG GAAGATGGCCGTACTCTTTCTGACTATGCGTCTGAGAGGTGGTATGCAGATCTTCGTGAAGACCCTGACCGGCAAGACCAATGT GAAGGCCAAGATCCAGGATAAAGAAGGCATCCCTCCCGACCAGCAGAGGGCACTCTTTCTGACTACAACATCCAGAAGGAGTCG ACCCTGCACCTGGTCCTGCTGACCGGCAAGACCATCACTCTGGAGGTGGAGCCCAGTGACACCATCGAAAATCCCGACCAGCAG…arrow_forward
- You plan to clone exon 11 of the HEXA gene (Section C) into the multiple cloning site (MCS) of the plasmid vector pUC19 illustrated below. You PCR amplify only the 184 bp DNA region representing exon 11 of HEXA from human DNA as a blunt-ended dsDNA fragment, and purify the amplicon. Next you digest the vector pUC19 with the restriction enzyme (RE) Smal to obtain linear plasmid DNA. You mix the PCR product and linear plasmid, add some DNA ligase enzyme in an appropriate buffer, and incubate overnight at 16°C. The ligation mixture is used to transform competent E. coli cells, which are subsequently streaked out onto agar plates containing ampicillin, X-gal and IPTG. HEXA exon 11 sequence: attcagccagacacaatcatacaggtgtggcgagaggatattccagtgaactatatgaaggagctggaactggtc accaaggccggcttccgggcccttctctctgccccctggtacctgaaccgtatatcctatggccctgactggaag gatttctacatagtggaacccctggcatttgaag PUC 19 plasmid map: 2686 1 Amp 0 lacZ EcoRI (390) Smal (410) BamHI (420) MCS Kpnl (430) LacR binding site Plac Pstl…arrow_forwardDescribe the process for shotgun sequencing of a genome. Practice aligning the two sets of sequenced fragments below, to determine the order of the fragments and the complete sequence.arrow_forwardA virus with a circular double-stranded DNA chromosome contains approximately 10,000 bp. You want to begin characterizing this chromosome by making a map of the cleavage sites of three restriction endonucleases: EcoRI, Hind III, and BamHI. You digest the viral DNA under conditions that allow the endonuclease reactions to go to completion and then subject the digested DNA to electrophoresis on agarose to determine the lengths of the restriction fragments produced in each reaction. Based on the resulting data, draw a map of the viral chromosome indicating the relative positions of the cleavage sites for these restriction endonucleases: Endonuclease EcoRI HindIII BamHI EcoRI + HindIII EcoRI + BamHI HindIII+ BamHI EcoRI + HindIII + BamHI Length of fragments (kb) 6.9, 3.1 5.1, 4.4, 0.5 10.0 3.6, 3.3, 1.5, 1.1, 0.5 5.1, 3.1, 1.8 4.4, 3.3, 1.8, 0.5 3.3, 1.8, 1.5, 1.1, 0.5arrow_forward
- Kpn I and Acc 65I are restriction enzymes that identify and cleave the same 6-bp sequence. The sticky end created by Kpn I cleavage, on the other hand, cannot be directly ligated to the sticky end formed by Acc 65I cleavage. Please explain why.arrow_forwardAfter restriction enzymes cut, they contain unpaired bases. Type II restriction enzymes leave ends that may be 5' overhanging, 3' overhanging, or blunt. In all cases each end is left with a 3' OH and a 5' phosphate. All blunt ends, and any complementary overhanging ends may be re-ligated with T4 DNA ligase, as long as at least one 5'- phosphate is present. In the tables below G^AATTC means that the end after cutting with enzyme will be: -----G 3' -----CTTAA 5' GTGCA^C means that the end will be: -----GTGCA 3' -----C 5' Which RE’s from table below have a 5’ overhang? Which ones have a 3’ Overhang? AccI GT^CGAC BamHI G^GATCC ClaI AT^CGAT NsiI ATGCA^T PstI CTGCA^G BglII A^GATCT TaqI T^CGAarrow_forwardThe restriction enzymes Kpn I and Acc 65I recognize and cleave the same 6-bp sequence. However, the sticky end formed from Kpn I cleavage cannot be ligated directly to the sticky end formed from Acc 65I cleavage. Explain why.arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





