
Biochemistry: Concepts and Connections (2nd Edition)
2nd Edition
ISBN: 9780134641621
Author: Dean R. Appling, Spencer J. Anthony-Cahill, Christopher K. Mathews
Publisher: PEARSON
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 6, Problem 23P
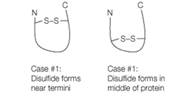
Disulfide bonds have been shown to stabilize proteins (i.e., make them less likely to unfold). Consider the cases shown schematically below for two variants of the same protein. In case #1 the disulfide forms between Cys residues that have been introduced near the protein N- and C-termini, and in case #2 the disulfide forms between Cys residues that have been introduced in the middle of the protein sequence. Which protein is likely to be more stable? (Note: Assume the disulfide bond is intact in both the unfolded and folded states). Explain your reasoning.
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
Match the three types of neurotransmitters to their relative size (largest to smallest):
Largest
Peptide neurotransmitter
✓
Second largest
[Choose]
Smallest
>
[Choose ]
[Choose ]
Amino acid neurotransmitter
Peptide neurotransmitter
Amine neurotransmitter
need help not sure why its wrong please help
Why the 2nd choice is correct?
Chapter 6 Solutions
Biochemistry: Concepts and Connections (2nd Edition)
Ch. 6 - Prob. 1PCh. 6 - Bovine pancreatic trypsin inhibitor (BPTI; Figure...Ch. 6 - A schematic structure of the subunit of...Ch. 6 - In the protein adenylate kinase, the C-terminal...Ch. 6 - Give two reasons to explain why a proline residue...Ch. 6 - Consider a small protein containing 101 amino acid...Ch. 6 - a. Based on a more conservative answer to Problem...Ch. 6 - The following sequence is part of a globular...Ch. 6 - a. A protein is found to be a tetramer of...Ch. 6 - Under physiological conditions, the protein...
Ch. 6 - Theoretical and experimental measurements show...Ch. 6 - The peptide hormone vasopressin is used in the...Ch. 6 - A protein gives under conditions of buffer...Ch. 6 - A protein gives a single band on SDS get...Ch. 6 - It has been postulated that the normal...Ch. 6 - Below are shown two views of the backbone...Ch. 6 - Do you expect a Pro Gly mutation in a...Ch. 6 - Rank the following in terms of predicted rates...Ch. 6 - Shown below are two cartoon views of the small...Ch. 6 - Prob. 20PCh. 6 - In most cases, mutations in the core of protein...Ch. 6 - A Leu Ala mutation at a site buried the core of...Ch. 6 - Disulfide bonds have been shown to stabilize...Ch. 6 - Cartoon renderings of the proteins Top 7 and adaH2...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biochemistry and related others by exploring similar questions and additional content below.Similar questions
- a) What are the differences between the Direct & Indirect Immunofluorescence Assays? (0.5 mark) b) What are the advantages of the Indirect Immunofluorescence Assays? (0.5 mark) c) A Super-Resolution Imaging Technique was developed in 2018 using imidazole, a His-tag ligand conjugated with a fluorophore to report the presence of a recombinant His-tag protein target, (Sci Rep, 2018, 8:5507). How does this technique improve the image quality? (2 marks)arrow_forwarda) What are the differences between the Direct & Indirect Immunofluorescence Assays? b) What are the advantages of the Indirect Immunofluorescence Assays? c) A Super-Resolution Imaging Technique was developed in 2018 using imidazole, a His-tag ligand conjugated with a fluorophore to report the presence of a recombinant His-tag protein target, (Sci Rep, 2018, 8:5507). How does this technique improve the image quality?arrow_forwardCalculate the number of ATP produced from oxidation of 1 molecule of glucosearrow_forward
- Example 1: 1. Suppose an enzyme (MW = 5,000 g/mole) has a concentration of 0.05 mg/L. If the kcat is 1 x 10 s, what is the theoretical maximum reaction velocity for the enzyme? A) 1050 µM/s. B) 100 µM/s. C) 150 μM/s. D) 105 μM/s.arrow_forwardIn 1956, E. P. Kennedy and S. B. Weiss published their study of membrane lipid phosphatidylcholine (lecithin) synthesis in rat liver. Their hypothesis was that phosphocholine joined with some cellular component to yield lecithin. In an earlier experiment, incubating 32 P-labeled phosphocholine at physiological temperature (37 °C) with broken cells from rat liver yielded labeled lecithin. This became their assay for the enzymes involved in lecithin synthesis. Determine the optimal pH for this enzyme and characterize the enzyme activity at different pH values. -O-P-O-CH2-CH₁₂-N(CH3)3 Phosphocholine H₂C-O-C-R HC-O-C-R2 + + + Cell fraction + ? HC-O-P-O-CH₁₂-CH₂-N(CH), O Phosphatidylcholine The researchers then centrifuged the broken cell preparation to separate the membranes from the soluble proteins. They tested three preparations: whole extract, membranes, and soluble proteins. Table 1 summarizes the results. Table 1: Cell fraction requirement for incorporation of 32p-phosphocholine into…arrow_forwardResearchers isolated an unknown substance, X, from rabbit muscle. They determined its structure from the following observations and experiments. (a) Qualitative analysis showed that X was composed entirely of C, H, and O. A weighed sample of X was completely oxidized and the H2O and CO2 produced were measured. This quantitative analysis revealed that X contained 40.00% C, 6.71% H, and 53.29% O by weight. (b) The molecular mass of X, as determined by mass spectrometry, was 90.00 atomic mass units (u). (c) Infrared spectroscopy showed that X contained one double bond. (d) X dissolved readily in water, and the solution demonstrated optical activity when tested in a polarimeter. (e) The aqueous solution of X is acidic. What is the empirical formula of X?arrow_forward
- Show work. don't give Ai generated solution....give correct solutionarrow_forwardBiochemistry What is the process of "transamination" in either the muscles or the liver, that involves keto acid or glutamic acid? Please explain how the steps work. Thank you!arrow_forwardBiochemistry Please help. Thank you What is the importance of glutamic acid in the metabolism of nitrogen from amino acids? (we know therole; it’s used to remove the nitrogen from amino acids so that the remaining carbon skeleton can bebroken down by the “usual” pathways, but what is the important, unique role that only glutamicacid/glutamate can do?)arrow_forward
- Biochemistry Please help. Thank you When carbamyl phosphate is joined to L-ornathine, where does the energy for the reaction come from?arrow_forwardBiochemistry Question Please help. Thank you What is the function of glutamate dehydrogenase?arrow_forwardBiochemistry Question Please help. Thank you How and why does a high protein diet affect the enzymes of the urea cycle?arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning
Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning Human Biology (MindTap Course List)BiologyISBN:9781305112100Author:Cecie Starr, Beverly McMillanPublisher:Cengage Learning
Human Biology (MindTap Course List)BiologyISBN:9781305112100Author:Cecie Starr, Beverly McMillanPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning

Human Heredity: Principles and Issues (MindTap Co...
Biology
ISBN:9781305251052
Author:Michael Cummings
Publisher:Cengage Learning

Biology: The Dynamic Science (MindTap Course List)
Biology
ISBN:9781305389892
Author:Peter J. Russell, Paul E. Hertz, Beverly McMillan
Publisher:Cengage Learning

Biology Today and Tomorrow without Physiology (Mi...
Biology
ISBN:9781305117396
Author:Cecie Starr, Christine Evers, Lisa Starr
Publisher:Cengage Learning

Biology (MindTap Course List)
Biology
ISBN:9781337392938
Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. Berg
Publisher:Cengage Learning

Human Biology (MindTap Course List)
Biology
ISBN:9781305112100
Author:Cecie Starr, Beverly McMillan
Publisher:Cengage Learning

Biochemistry
Biochemistry
ISBN:9781305577206
Author:Reginald H. Garrett, Charles M. Grisham
Publisher:Cengage Learning
Biomolecules - Protein - Amino acids; Author: Tutorials Point (India) Ltd.;https://www.youtube.com/watch?v=ySNVPDHJ0ek;License: Standard YouTube License, CC-BY