
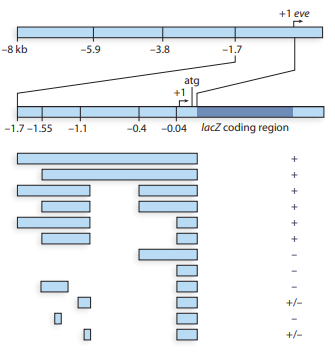
The Drosophila even
Want to see the full answer?
Check out a sample textbook solution
Chapter 16 Solutions
Genetic Analysis: An Integrated Approach (2nd Edition)
- The MAT locus allows yeast to switch mating type through a very complex mechanism. However, it has informed us a great deal about what aspects of gene expression typical to all organisms? Options: higher order changes in chromatin affect transcriptional efficiency that general transcription factors must first bind directly to histone tails and only then can they interact with their cognate binding sites that DNA methylation is involved in this silencing mechanism that SIR2 is required for all types of transcriptional repression that expression of Pol III genes provides a means of identifying active chromatinarrow_forwardThe UG4 gene is expressed in the stem and leaf tissue of the plant Arabidopsis thaliana. To identify potential mechanisms regulating UG4 gene expression, six small deletion mutations are made in cloned sequence containing the upstream regulatory region. The full length mutant segments Which mutation(s) affect an enhancer? Why? Which mutations identify the promoter? Why? Speculate about the reason for different transcription rated obtained for fusion constructs E and F.arrow_forwardName the lambda promoters whose expression is regulated by the cro protein. For each promoter you named, is cro an activator or a repressor of transcription from that promoter?arrow_forward
- The IMD2 promoter contains three upstream transcription start sites (TSS) that are utilized under high GTP conditions and a single downstream TSS (-106) that is normally only utilized under low GTP conditions. In a wild type cell, expression of IMD2 mRNA only occurs if transcription initiates from the -106 TSS. In 300 words or less, describe: 1.) The normal function of Ssl2, and 2.) why a mutation in Ssl2, that increases its catalytic rate, would allow expression of the IMD2 ORF under high GTP conditions. (Conditions under which the IMD2 ORF is NOT expressed in the wild type.)arrow_forwardGR and PPAR are transcription factors that bind to GRE and PPARE sequences respectively and activate transcription of genes. A reporter cell line is created in which the the green fluoresecent protein (GFP) is controlled by a GRE sequence and the pink fluorescent protein mCherry is under control of a PPARE sequence. If the gene for GR is introduced into the reporter cell line, the cells produce a green color. Chimeric proteins are created in which the DNA Binding Domains (DBD) and Activation Domains (AD) of the transcription factors are introduced into various cell lines. Match the following cell-types with the fluorescent color(s) you would expect the cells to produce.arrow_forwardGal4 is a transcription factor that activates transcription of galactose metabolism genes in yeast. These genes are ‘turned on’ when yeast cells need to metabolize galactose. To identify promoter sequences necessary for regulation of transcription of GAL1, reporter gene fusions were made and introduced into yeast cells. Deletions of GAL1 promoter were cloned upstream of LacZ gene. β-Galactosidase activity was measured in presence of galactose. Shown below is a representation of the results obtained. In the diagrams below (not to scale!): • Construct 1 contains ~ 130bp of the promoter, which is predicted to have all the predicted/putative proximal promoter elements (indicated by the solid boxes) needed to regulate transcription of GAL1.• The stippled box is the core promoter.• The arrow represents the transcriptional start site for the reporter gene Lac Z• Number of + signs represents level of transcription• Star represents a mutation in DNA sequence at that location (few nucleotides…arrow_forward
- Based on the information in Figure 12-6, how doesGal4 regulate four different GAL genes at the sametime? Contrast this mechanism with how the Lac repressor controls the expression of three genes.arrow_forwardThe extracellular protein factor Decapentaplegic(Dpp) is critical for proper wing development in Drosoph-ila (Figure Q21–3A). It is normally expressed in a narrowstripe in the middle of the wing, along the anterior–pos-terior boundary. Flies that are defective for Dpp formstunted “wings” (Figure Q21–3B). If an additional copyof the gene is placed under control of a promoter that isactive in the anterior part of the wing, or in the posterior part of the wing, a large mass of wing tissue composed ofnormal-looking cells is produced at the site of Dpp expres-sion (Figure Q21–3C and D). Does Dpp stimulate cell divi-sion, cell growth, or both? How can you tell?arrow_forwardA protein or RNA that regulates gene expressionin trans—either at the level of transcription, RNAsplicing, or translation—must have specificity for onetarget gene or a group of target genes. Explain howspecificity is achieved in each casearrow_forward
- An electrophoretic mobility shift assay can be used to study the binding of proteins to a segment of DNA. In the results shown here, an EMSA was used to examine the requirements for the binding of RNA polymerase |l (from eukaryotic cells) to the promoter of a protein-encoding gene. The assembly of general transcription factors and RNA polymerase Il at the core promoter is described in Week 4. In this experiment, the segment of DNA containing a promoter sequence was 1100 bp in length. The fragment was mixed with various combinations of proteins and then subjected to an EMSA. Lane 1: No proteins added Lane 2: TFIID Lane 3: TFIIB Lane 4: RNA polymerase IIl Lane 5: TFIID + TFIIB Lane 6: TFIID + RNA 1 2 3 4 5 6. 7 polymerase II Lane 7: TFIID + TFIIB + RNA polymerase Il 1100 bp Explain the results.arrow_forwardA strain of Arabidopsis thaliana possesses a mutation in the APETALA2 gene. As a result of this mutation, much of the 3′ UTR of the mRNA transcribed from the gene is deleted. What is the most likely effect of this mutation on the expression of the APETALA2 gene?arrow_forwardMost eukaryotic promoters have binding sites for several different transcription factors, and the frequency with which transcription is initiated at a promoter depends on the specific combination of transcriptional regulators bound to their binding sites in that promoter. Transcription of the slither gene in garter snakes is regulated by the transcriptional activators Python and Boa and the transcriptional repressor Sidewinder. Each of these proteins has one binding site in the slither promoter; the affinity of Boa for its binding site is 30 times higher than the affinity of Python for its binding site and 6 times higher than the affinity of Sidewinder for its binding site. Under conditions where Sidewinder is 10 times more abundant than Python, and Python is 3 times as abundant as Boa, would you expect transcription of the slither gene to be activated or repressed? Show or briefly explain how/why you predicted the outcome you chose.arrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
