
Concept explainers
To estimate: The K value from the given data that describes the ligand binding to a protein.
Concept introduction: Dissociation constant is the equilibrium constant at which the ligand dissociates from the protein. It is denoted as K. Y is fractional saturation of the ligand occupying the ligand-binding site in the protein, or it can be defined as the number of ligands that has dissociated from the protein. K is achieved when fractional saturation is half at a particular ligand concentration.
Answer to Problem 1E
Answer: The dissociation constant is estimated by plotting the values in a graph and finding the ligand concentration when Y is 0.5 (half of fractional saturation).
Pictorial representation:
Fig.1 shows a graph with ligand (mM) in the X-axis and Y in the Y-axis.
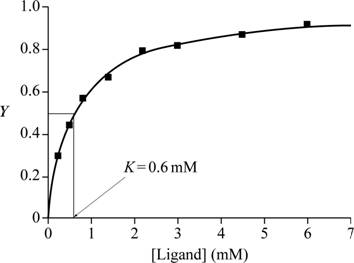
Fig.1: Graph of ligand (mM) vs Y to estimate K.
Explanation of Solution
Explanation:
When fractional saturation is half, it means that the ligand has occupied half of the ligand binding site in the protein. When fractional saturation is half, half of the ligand is bound to half the number of proteins in the medium. Half of proteins and half of ligands are free in the medium. This can be called as the equilibrium state of the ligand-protein binding reaction, and thus K is obtained at half of the fractional saturation.
A graph is plotted with ligand concentration in mM on the X-axis and Y, the fractional saturation, on the Y-axis with the given data. At half of fractional saturation, that is, at equilibrium state, the corresponding ligand concentration is 0.6 Mm, which is evident from the graph.
Conclusion: From the graph plotted for the given values, the K value is 0.6mM.
Want to see more full solutions like this?
Chapter 7 Solutions
Biochemistry 410/411 Textbook - 5th Edition - Custom Texas A&M University
- answer the questions and the example steps should be from carbohydrates glycolysis and citric acid cycle. Please put down reactions and structuresarrow_forwardidentify the general type of reaction catalyzed and an example step from glycolisis structure for each of the following enzymes/ co factor Kinase, isomerase, mutase, dehydrogenase, NAD+ , FADarrow_forwardfill in the blanks with the missing structures and give namesarrow_forward
- fill in the table and identify the general type of reaction catalayzed and an example step from the structures in the second page so you will answer the questions from the first page the second one is just a reference urgently!arrow_forwardPlease draw out the molecular structures of each molecule and show how each enzyme + cofactor would affect the following molecule in the human metabolic pathway. (This is a metabolic map)arrow_forwardPlease draw out the molecular structures of each molecule and show how an enzyme + cofactor would affect the following molecule in the human metabolic pathway to create energy.arrow_forward
- Please draw out the molecular structures of each molecule and show how each enzyme + cofactor would affect the following molecule in the human metabolic pathway.arrow_forwardPlease draw out the mechanism with curved arrows showing electron flow. Pyruvate is accepted into the TCA cycle by a “feeder” reaction using the pyruvate dehydrogenase complex, resulting in acetyl-CoA and CO2. Provide the mechanism for this reaction utilizing the TPP cofactor. Include the roles of all cofactors.arrow_forwardPyruvate is accepted into the TCA cycle by a “feeder” reaction using the pyruvate dehydrogenase complex, resulting in acetyl-CoA and CO2. Provide the mechanism for this reaction utilizing the TPP cofactor. Include the roles of all cofactors.arrow_forward
- The mitochondrial ATP synthase has 10 copies of the F0 subunit “c”, and the [H ] in the mitochondrial inner membrane space (IMS) is 6.31 x 10-8 M and the [H + ] in the matrix is 3.16 x 10-9 M. Calculate the minimum membrane potential (∆Ψ) necessary to make ATP synthesis thermodynamically favorable. [Assume ∆G' ofphosphate hydrolysis of ATP is - 45 kJ/mol.]arrow_forwardB- Vitamins are converted readily into important metabolic cofactors. Deficiency in any one of them has serious side effects. a. The disease beriberi results from a vitamin B 1 (Thiamine) deficiency and is characterized by cardiac and neurological symptoms. One key diagnostic for this disease is an increased level of pyruvate and α-ketoglutarate in the bloodstream. How does this vitamin deficiency lead to increased serumlevels of these factors? b. What would you expect the effect on the TCA intermediates for a patient suffering from vitamin B 5 deficiency? c. What would you expect the effect on the TCA intermediates for a patientsuffering from vitamin B 2 /B 3 deficiency?arrow_forwardPyruvate is accepted into the TCA cycle by a “feeder” reaction using the pyruvate dehydrogenase complex, resulting in acetyl-CoA and CO2. Provide a full mechanism for this reaction utilizing the TPP cofactor. Include the roles of all cofactors.arrow_forward
 BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman
BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman
Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY
Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON
Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON





