
Concept explainers
SCIENTIFIC INQUIRY
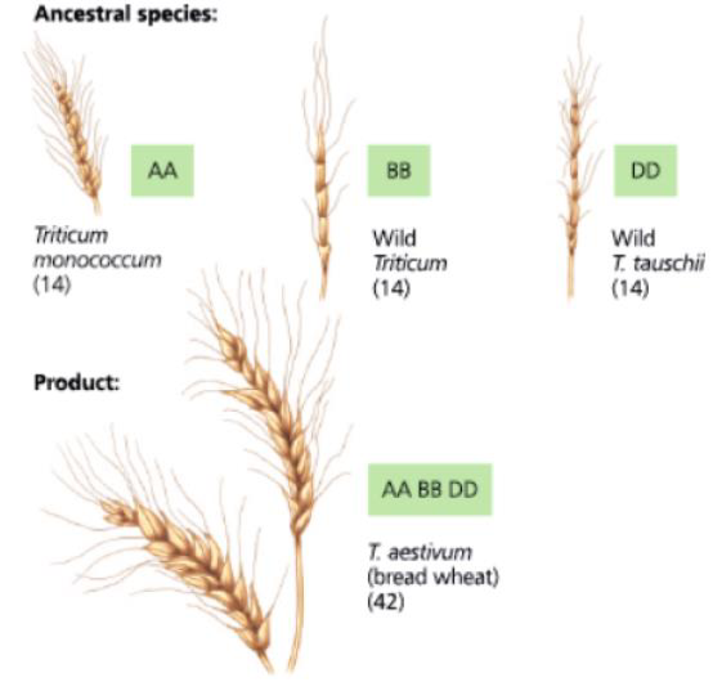
DRAW IT In this chapter, you read that bread wheat (Triticum aestivum) is an allohexaploid, containing two sets of chromosomes from each of three different parent species. Genetic analysis suggests that the three species pictured following this question each contributed chromosome sets to T. aestivum. (The capital letters here represent sets of chromosomes rather than individual genes.) Evidence also indicates that the first polyploidy event was a spontaneous hybridization of the early cultivated wheat species T. monococcum and a wild Triticum grass species. Based on this information, draw a diagram of one possible chain of events that could have produced the allohexaploid T. aestivum.
Want to see the full answer?
Check out a sample textbook solution
Chapter 22 Solutions
Campbell Biology in Focus (2nd Edition)
Additional Science Textbook Solutions
Physical Science
Biological Science (6th Edition)
Campbell Essential Biology (7th Edition)
HUMAN ANATOMY
Laboratory Manual For Human Anatomy & Physiology
- Does it show the level of proteins? What about the amount? Levels of protein activation? How can you tell? Does the thickness tell you anything? What about the number of the lines? And the other questionsarrow_forwardKD 200- 116- 66- Vec ATF6 (670) ATF6 (402) ATF6 (373) ATF6 (366) I I 45- 1 2 3 4 5 ATFG (360) (e/c) 9V ATFG (402) g ant- ATF anti-KDEL DAPI barrow_forwardWestern blot results: what information can you get? Presence of proteins of your interest Levels of protein expression Levels of protein activation (must use activation state-specific antibody) Decreased function of the ATM kinase in aging mice. A C57BL/6 female 6 month Con IR 20 month C57BL/6 male 6 month 28 month Con IR Con IR Con IR p-ATM (S1981) ATM P-p53 (ser18) Actinarrow_forward
- Does it show the level of proteins? What about the amount? Levels of protein activation? How can you tell? Does the thickness tell you anything? What about the number of the lines?arrow_forwardWB: Protein of interest visualized by fluorescent Protein A Protein Barrow_forwardQuestion #4: Assume you are able to use CRISPR to create an allele that will convert a cross-pollinated, sexually reproducing crop plant into an obligate apomict. Your edited obligate apomict plants retains all the CRISPR "machinery" necessary to convert the "sexually reproducing" allele to the "obligate apomict" allele. You plant 100 hectares of your edited obligate apomicts in order to increase seed for sale the following year. Neighboring farms and seed producers are growing many different un-edited sexually reproducing varieties of the crop. If your neighbors plant seed harvested from their crops that was pollinated by your crop, should they expect these seeds to generate apomictic or non-apomictic plants? Type your answer here:arrow_forward
- calculate the questions showing the solution including variables,unit and equations all the questiosn below using the data.show solving and answer a) B1, b) B2, c) hybrid rate constant (1) d) hybrid rate constant (2) e) t1/2,dist t1/2,absorb f) t1/2,elim k) apparent central compartment volume (V1,app) p) total AUC (using short cut method) apparent volume of distribution based on AUC (VAUC,app) apparent clearance (CLapp) absolute bioavailabilty of oral route ( AUCiv =116ml)arrow_forwardPlease help me to draw this by hand. In as much detail as possible, hand draw a schematic diagram of the hypothalamic-pituitary-gonad (HPG) axis in the human female. Be sure to include all the relevant structures and hormones. You must define all abbreviations the first time you use them. Please include (and explain) the feedback loops.arrow_forwardPlease refer belowarrow_forward
- AaBbCc X AaBbCc individuals are crossed. What is the probability of their offspring having a genotype AABBCC?arrow_forwardcircle a nucleotide in the imagearrow_forward"One of the symmetry breaking events in mouse gastrulation requires the amplification of Nodal on the side of the embryo opposite to the Anterior Visceral Endoderm (AVE). Describe one way by which Nodal gets amplified in this region." My understanding of this is that there are a few ways nodal is amplified though I'm not sure if this is specifically occurs on the opposite side of the AVE. 1. pronodal cleaved by protease -> active nodal 2. Nodal -> BMP4 -> Wnt-> nodal 3. Nodal-> Nodal, Fox1 binding site 4. BMP4 on outside-> nodal Are all of these occuring opposite to AVE?arrow_forward
 Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax





