
Concept explainers
(a)
To describe: The chemical transformation which is required to convert glutamate to (2S)-4-amino-2-hydroxybutyrate (AHBA).
Introduction:
Glutamate comes under the category of non-essential amino acid and it contains two carboxyl groups. It is an acidic amino acid which is important part of the neurotransmission process. AHBA (4-amino-2-bydoxy butyrate) it is main component of many antibiotics.
(a)
Explanation of Solution
Pictorial representation: Fig.1 shows formation of AHBA from glutamate.
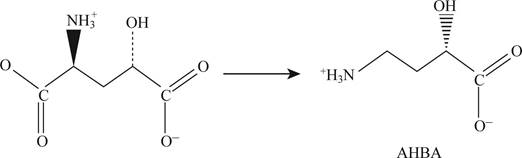
Fig.1: Formation of AHBA from glutamate.
Explanation:
AHBA (4-amino-2-bydoxy butyrate) is formed from the glutamate by the process of decarboxylation. When there is a removal of α-carboxyl group from the glutamate and addition of one –OH group to the γ-carbon then AHBA is formed which is given in Fig. 1.
(b)
To determine: How Btrl can act as an acyl carrier protein with a Coenzyme A prosthetic group.
Introduction:
Coenzyme A is important in the synthesis and oxidation of the fatty acids and it also help in the citric acid cycle during the oxidation of the pyruvate.
(b)
Explanation of Solution
Explanation:
Btrl and acyl carrier proteins have similarities in terms of their sequences. When this compound incubated in CoA conditions it increases its molecular weight. It is because CoA molecule can bind to Btrl molecule when CoA is added to the serine residue (any one) with the replacement of the hydroxyl group with a 4’-phosphopantertheine group
The molecular weight of the CoA is 17;
And the molecular weight of 4’phosphopantetheine is 356;
Molecular mass of purified Btrl protein is 11,812.
So, the overall molecular weight of the Btrl which could act as an acyl carrier protein with a Coenzyme A would be as follows:
This calculated molecular weight is very much close to the observed molecular weight 12,153.
Conclusion:
Btrl can act as an acyl carrier protein with a Coenzyme A because the expected molecular weight is very close to the calculated value of molecular weight which is 12,153.
(c)
To determine: The structures which are consistent with the proposed structure of γ-glutamyl-S-Btrl.
Introduction:
Alpha carboxyl group is any organic molecule group that contains basic amino group and acidic carboxyl group which is very unique to each amino acid.
(c)
Explanation of Solution
Pictorial representation: Fig. 2 shows glutamyl-S-Btrl.
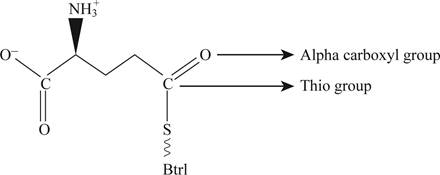
Fig.2: Glutamyl-S-Btrl
Explanation:
One more structure which is consistent is that a thio-ester could formed with the α-carboxyl group which is represented in the Fig. 2. Thio-ester is the compound which has
(d)
To determine: The chemical basis of the γ-glutamyl-S-Btrl can be correct because the alpha carboxyl group might be removed at some stage in the process.
Introduction:
Organic molecule group such as alpha carboxyl group that contains basic amino group and acidic carboxyl group and it is very unique to each amino acid.
(d)
Explanation of Solution
Explanation:
The process of removal of a carboxyl group from a compound is termed as decarboxylation. In many reactions, carboxyl group should be in a free state for an amino acid to be removed. It is very difficult to imagine that decarboxylation process can occur with a carboxyl group in its thio-ester form.
(e)
To determine: The most likely structure of the BtrK species
Introduction:
Decarboxylation is a
(e)
Explanation of Solution
Pictorial representation: Fig 3 shows most likely structure of the BtrK
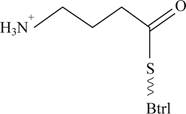
Fig 3: Most likely structure of the BtrK
Explanation:
The molecular species of Mr is 12,240 when the γ-glutymyl-S-Btrl is incubated with purified Btrl.
The molecular species of Mr is 12,281 of γ-glutymyl-S-Btrl.
Therefore,
Due to decarboxylation of the BtrK, carboxyl group would be deleted from the structure and only
(f)
To determine: The molecular structure of the species whose molecular mass is 12,370.
Introduction:
A pure substance is made from molecules with the average geometrical structure. The chemical formula and the structure of a molecule are the two important factors that determine.
(f)
Explanation of Solution
Pictorial representations: Fig 4 shows expected molecular structure of the species whose molecular mass is 12,370.
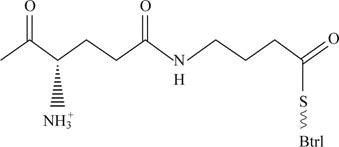
Fig 4: Expected molecular structure of the species whose molecular mass is 12,370.
Explanation:
The Molecular structure of the species whose molecular mass is 12,370 is represented in the figure 4. The molecular species of Mr is 12,240 when the γ-glutymyl-S-Btrl is incubated with purified Btrl. The Molecular structure of the species whose molecular mass is 12,370.
Therefore,
This molecular structure would have free amino group and alpha carboxyl group and thio group will remain in the new species.
(g)
To propose: The plausible pathway for the synthesis of AHBA with the help of enzymes that catalyze each step and any kind of cofactors and substrate required during the pathway.
Introduction:
In the plausible pathways different kind of enzyme and cofactor and substrate are used with the help of these factors a useful product can be synthesized.
(g)
Explanation of Solution
Pictorial representation: Fig.5 shows the plausible pathway for t.he synthesis of AHBA
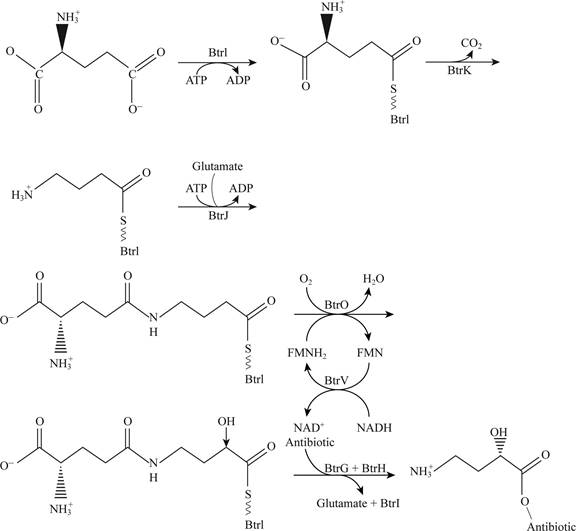
Fig.5: The plausible pathway for the synthesis of AHBA
Explanation:
The plausible pathway for the synthesis of AHBA and respective addition of antibiotics is represented in the Fig. 5. This pathway includes the involvement of co-factor such as ATP, NAD and glutamate amino acid.
Want to see more full solutions like this?
Chapter 22 Solutions
SAPLINGPLUS FOR PRINCIPLES OF BIOCHEMIS
- Sodium fluoroacetate (FCH 2CO2Na) is a very toxic molecule that is used as rodentpoison. It is converted enzymatically to fluoroacetyl-CoA and is utilized by citratesynthase to generate (2R,3S)-fluorocitrate. The release of this product is a potentinhibitor of the next enzyme in the TCA cycle. Show the mechanism for theproduction of fluorocitrate and explain how this molecule acts as a competitiveinhibitor. Predict the effect on the concentrations of TCA intermediates.arrow_forwardIndicate for the reactions below which type of enzyme and cofactor(s) (if any) wouldbe required to catalyze each reaction shown. 1) Fru-6-P + Ery-4-P <--> GAP + Sed-7-P2) Fru-6-P + Pi <--> Fru-1,6-BP + H2O3) GTP + ADP <--> GDP + ATP4) Sed-7-P + GAP <--> Rib-5-P + Xyl-5-P5) Oxaloacetate + GTP ---> PEP + GDP + CO 26) DHAP + Ery-4-P <--> Sed-1,7-BP + H 2O7) Pyruvate + ATP + HCO3- ---> Oxaloacetate + ADP + Piarrow_forwardTPP is also utilized in transketolase reactions in the PPP. Give a mechanism for theTPP-dependent reaction between Xylulose-5-phosphate and Ribose-5-Phosphate toyield Glyceraldehyde-3-phosphate and Sedoheptulose-7-Phosphate.arrow_forward
- What is the difference between a ‘synthetase’ and a ‘synthase’?arrow_forwardIn three separate experiments, pyruvate labeled with 13C at C-1, C-2, or C-3 is introduced to cells undergoing active metabolism. Trace the fate of each carbon through the TCA cycle and show when each of these carbons produces 13CO2.a. Glucose is similarly labeled at C-2 with 13C. During which reaction will this labeled carbon be released as 13CO2?arrow_forwardDraw the Krebs Cycle and show the entry points for the amino acids Alanine,Glutamic Acid, Asparagine, and Valine into the Krebs Cycle. How many rounds of Krebs will be required to waste all Carbons of Glutamic Acidas CO2?arrow_forward
- Suppose the data below are obtained for an enzyme catalyzed reaction with and without the inhibitor I. (s)( mM) 0.2 0.4 0.8 1.0 2.0 4.0 V without i (mM/min) 5.0 7.5 10.0 10.7 12.5 13.6 V with I (mM/min) 3.0 5.0 7.5 8.3 10.7 12.5 Make a Lineweaver Burke plot for this data using graph paper or a spreadsheet Calculate KM and Vmax without inhibitor. What type of inhibition is observed? show graph and work 2. Give the Lineweaver Burk equation and define all the parameters. 3. When substrate concentration is much greater than Km, the rate of catalysis is almost equal to a. kcat b. none of these c. all of these d. Kd e. Vmaxarrow_forwardPlease explain the process of how an axon degenerates in the central nervous system following injury and how it affects the neuron/cell body, as well as presynaptic and postsynaptic neurons. Explain processes such as chromatolysis and how neurotrophin signaling works.arrow_forwardPlease help determine the Relative Response Ratio of my GC-MS laboratory: Laboratory: Alcohol Content in Hand Sanditizers Internal Standard: Butanol Standards of Alcohols: Methanol, Ethanol, Isopropyl, n-Propanol, Butanol Recorded Retention Times: 0.645, 0.692, 0.737, 0.853, 0.977 Formula: [ (Aanalyte / Canalyte) / (AIS / CIS) ]arrow_forward
- Please help determine the Relative Response Ratio of my GC-MS laboratory: Laboratory: Alcohol Content in Hand Sanditizers Internal Standard: Butanol Standards of Alcohols: Methanol, Ethanol, Isopropyl, n-Propanol, Butanol Recorded Retention Times: 0.645, 0.692, 0.737, 0.853, 0.977 Formula: [ (Aanalyte / Canalyte) / (AIS / CIS) ]arrow_forwardplease draw it for me and tell me where i need to modify the structurearrow_forwardPlease help determine the standard curve for my Kinase Activity in Excel Spreadsheet. Link: https://mnscu-my.sharepoint.com/personal/vi2163ss_go_minnstate_edu/_layouts/15/Doc.aspx?sourcedoc=%7B958f5aee-aabd-45d7-9f7e-380002892ee0%7D&action=default&slrid=9b178ea1-b025-8000-6e3f-1cbfb0aaef90&originalPath=aHR0cHM6Ly9tbnNjdS1teS5zaGFyZXBvaW50LmNvbS86eDovZy9wZXJzb25hbC92aTIxNjNzc19nb19taW5uc3RhdGVfZWR1L0VlNWFqNVc5cXRkRm4zNDRBQUtKTHVBQldtcEtWSUdNVmtJMkoxQzl3dmtPVlE_cnRpbWU9eEE2X291ZHIzVWc&CID=e2126631-9922-4cc5-b5d3-54c7007a756f&_SRM=0:G:93 Determine the amount of VRK1 is present 1. Average the data and calculate the mean absorbance for each concentration/dilution (Please over look for Corrections) 2. Blank Correction à Subtract 0 ug/mL blank absorbance from all readings (Please over look for Corrections) 3. Plot the Standard Curve (Please over look for Corrections) 4. Convert VRK1 concentration from ug/mL to g/L 5. Use the molar mass of VRK1 to convert to M and uM…arrow_forward
 BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman
BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman
Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY
Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON
Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON





