
Concept explainers
Answers to all problems are at the end οΓthis book. Detailed solutions are available in the Student Solutions Manual. Study Guide, and Problems Book.
Characterizing a Covalent Enzyme Inhibitor Tosyl-L-phenylalanme cfaloromethyl
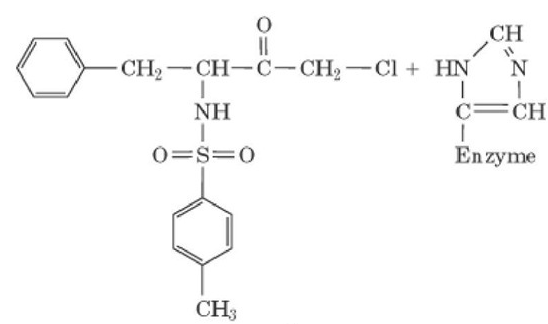
- Propose a mechanism for the inactivation reaction, indicating the structure of the produce(s).
- State why this inhibitor is specific tor cJiymotrypsin.
- Propose a reagent based on the structure of TPCK that might be an effective inhibitor of trypsin.
a.
To propose: The mechanism for the inactivation reaction which indicates the structure of the products.
Introduction:
Covalent enzyme inhibitor like TPCK is slowly reversible or totally irreversible. In many cases, these inhibitors are used to validate the pathway or target. There are targeted covalent enzyme inhibitors that have a slow offset, high binding efficiency as well as high potency to bind, increased selectivity decreased propensity for target-based drug resistance and prolonged pharmacodynamic effect.
Explanation of Solution
Chymotrypsin undergoes covalent inactivation reaction by nucleophilic attack on the
So, the mechanism of inactivation reaction is given below:

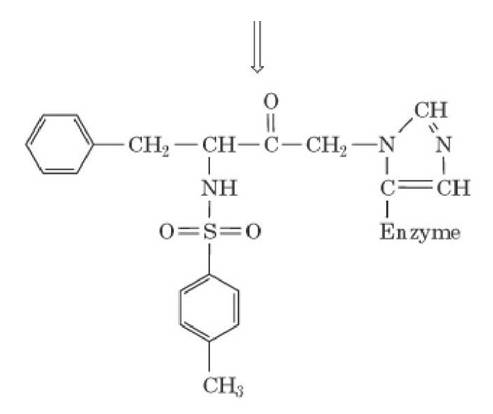
Hence, from the above discussion, it is clear that chymotrypsin undergoes covalent inhibition reaction.
b.
To propose: The reason for this inhibitor to be specific for chymotrypsin.
Introduction:
Covalent enzyme inhibitor like TPCK is slowly reversible or totally irreversible. In many cases, these inhibitors are used to validate the pathway or target. There are targeted covalent enzyme inhibitors that have a slow offset, high binding efficiency as well as high potency to bind, increased selectivity decreased propensity for target-based drug resistance and prolonged pharmacodynamic effects. Chymotrypsin plays an important part in the breakdown of proteins which is called proteolysis. The pancreas synthesizes the chymotrypsin. It cleaves the peptide amino bonds to digest the proteins.
Explanation of Solution
The inhibitor is chymotrypsin specific because the chloromethyl keto group reaches the active site by the hydrophobic reactions of chymotrypsin and the phenyl group of TPCK.
Hence, from the above discussion, it is clear that the inhibitor is chymotrypsin specific.
c.
To identify: A reagent that can be an effective inhibitor of Trypsin.
Introduction:
Chymotrypsin plays an important part in the breakdown of proteins which is called proteolysis. The pancreas synthesizes the chymotrypsin. It cleaves the peptide amino bonds to digest the proteins.
Explanation of Solution
The phenylalanine residue of TPCK has to be replaced with lysine or arginine to produce the specific reagents for the enzyme Trypsin.
Hence, from the above discussion, it is clear that the residual portion of TPCK only produces the specific reagent for Trypsin.
Want to see more full solutions like this?
Chapter 14 Solutions
Biochemistry
- Characterize each term or phrase as pertaining to simple or facilitated diffusion. Simple diffusion Facilitated diffusion Answer Bank requires an input of free energy lipophilic molecules directly through membrane via channels polar molecules Na+arrow_forwardSort the descriptions into properties that describe either saturated phospholipids or unsaturated phospholipids. Saturated phospholipids Saturated and unsaturated phospholipids Unsaturated phospholipids Answer Bank have no double bonds in the fatty acid carbon chains have straight fatty acid tails have at least one double bond in the fatty acid tails have bent fatty acid tails are built upon a glycerol backbone make the membrane somewhat rigid at low temperatures allow the membrane to remain fluid and flexible at low temperatures fatty acid tails pack tightly together maintain some space between adjacent phospholipidsarrow_forwardPlace the events of an action potential in order, starting and ending with a cell at its resting membrane potential. Cell starts at its resting membrane potential. Cell returns to its resting membrane potential. Answer Bank K+ channels fully open, and Na+ channels are inactivated. K* rushes out of the cell, causing repolarization. K+ channels close slowly, resulting in hyperpolarization. Na+ channel gates reset. Fast Na+ and slow K+ channels are activated. Na rushes into the cell, causing membrane depolarization. Ligand activation of the acetylcholine receptor depolarizes the membrane.arrow_forward
- Glucose and fructose are reducing sugars. Sucrose, or table sugar, is a disaccharide consisting of both fructose and glucose. Is sucrose a reducing sugar? Why or why not? No, because only one anomeric carbon is involved in the glycosidic linkage. No, because both anomeric carbons are involved in the glycosidic linkage. Yes, because the fructose unit can convert to the open-chain form. Yes, because the glucose unit can convert to the open-chain form. Which statements about reducing sugars are true? The oxidation of a reducing sugar forms a carboxylic acid sugar. D-Arabinose (an aldose) is a reducing sugar. Reducing sugars contain keto groups instead of aldehyde groups. A disaccharide with its anomeric carbons joined by the glycosidic linkage cannot be a reducing sugar. A reducing sugar will not react with the Cu² + in Fehlings's reagent.arrow_forwardExamine the pairs of molecules and identify the more-reduced molecule in each pair. H-C- CH, OH CH HO-C-H CH₁₂ Pyruvate Ethanol Acetaldehyde Lactate COO H-C H H- -C-H COO- Succinate Fumarate -OOC COO H COO- H――000- CH₂ COO- Oxalosuccinate H-C-OH OOC-C-H CH₂ COO Isocitratearrow_forwardClassify each description as characterizing facilitated diffusion, primary active transport, secondary active transport, or both primary and secondary active transport. Facilitated diffusion Primary active transport Secondary active transport Primary and secondary active transport Answer Bank requires ATP includes lactose permease directly uses ATP hydrolysis to pump substances across the membrane includes the Na+-K+ ATPase pump always moves more than one substance at a time movement of substances against an electrochemical gradient does not require energy input includes uniporters uses energy stored in electrochemical gradients generated by pumpsarrow_forward
- Creatine is a popular dietary supplement. What is the biochemical rationale for the use of creatine? It would directly serve as an electron carrier to support the oxidation of fuel molecules and thus energy production. It would serve as an electron donor to support reductive biosyntheses required to sustain cellular function. It would be converted into creatine phosphate and thus serve as a rapid means of replenishing ATP during muscle contraction. It would promote the movement of ions through ion channels and thus power the synthesis of ATP during exercise. What type of exercise would benefit most from creatine supplementation? a leisurely walk sprinting yoga a long-distance runarrow_forwardAssign each statement to the corresponding polysaccharide. Chitin Starch Glycogen Cellulose Answer Bank is abundant in muscle and liver provides structural support for plants is the storage form of glucose in animals provides structural support for animals such as arthropods is a storage form of fuel in plant cells consists of N-acetylglucosamine residues comes in two forms: amylose and amylopectinarrow_forwardMatch each term with its description. has the molecular formula of (CHO), monosaccharides that differ at a single asymmetric carbon atom the storage form of glucose in animals the storage form of glucose in plants glycoprotein containing glycosaminoglycans the most abundant organic molecule in the biosphere N-acetylgalactosamine is a key component of this glycoprotein carbohydrate-binding proteins enzymes that synthesize oligosaccharides stereoisomers that are mirror images of each other Answer Bank lectins epimers starch mucoprotein carbohydrates glycogen glycosyltransferases cellulose enantiomers proteoglycanarrow_forward
- Complete the sentences describing membrane lipids by moving the names of the lipids to the appropriate sentence. Some lipids will be used more than once, and some sentences will require you to place two or three lipids. Answer Bank include two fatty acids joined to glycerol by ester linkages. do not contain glycerol. is a steroid. contain a sphingosine backbone. contain one or more sugars. usually have branched alkyl chains. glycolipids phosphoglycerides sphingomyelin cholesterol archacal lipidsarrow_forwardThe protein content of most plasma membranes is, on average, about 50% by weight. Myelin has a protein content of about 18%, whereas the internal membranes of mitochondria and chloroplasts may be composed of 75% protein. Label the membrane proteins on the diagram. Answer Bank lipid-anchored protein peripheral membrane protein integral membrane proteinarrow_forwardWhy don’t we see amino acids with certain properties (e.g., a straight-chain side chain with two carbons, multiple hydroxyl groups, or other unusual structures)?arrow_forward
 BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
