
Concept explainers
Answers to all problems are at the end of this book.. Detailed solutions are available in the Student Solutions Manual, Study Guide, and Problems Book.
Naming Sugars in the (R,S) System Use the inform tit ion in the Critical Developments in Biochemistry box titled "Rules for Description (if Chiral Centers in the (R.S) System" (Chapter 4) to name D-galactose using (R.S) nomenclature. Do the same for L-altrone.
Interpretation:
The name of
Concept introduction:
A carbon atom that has four non-equivalent atoms or groups attached to it is known as chiral carbon atom. Chiral carbon centers are also called as asymmetric or stereogenic centers.
The naming of chiral center and geometric isomers are based on Cahn-Ingold-Prelog priority rules. If the priority assigned to each group attached to the chirality center in a molecule is in a clockwise direction, then it is the R-stereoisomer, and if this is counter-clockwise, then it is the S-stereoisomer. R and S-stereoisomer are mirror images of each other.
Answer to Problem 7P
The name of
Explanation of Solution
The highest priority group in case of a monosaccharide is hydroxyl
The Fischer projection structure for
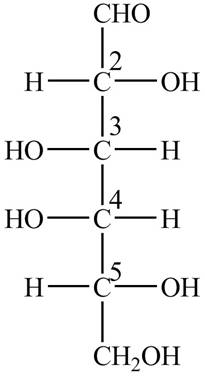
Figure 1
The substituents which extend to the side of
Thus, in the structure of
At
At
At
Thus, the name of
The Fischer projection structure for
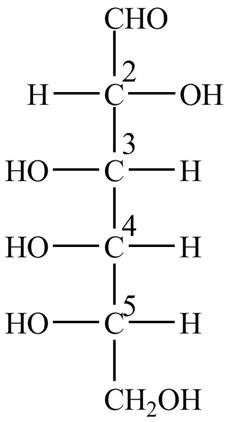
Figure 2
The substituents which extend to the side of
Thus, in the structure of
At
At
At
Thus, the name of
The name of
Want to see more full solutions like this?
Chapter 7 Solutions
EBK BIOCHEMISTRY
- The reduced coenzymes generated by the citric acid cycle donate electrons in a series of reactions called the electron-transport chain. The energy from the electron-transport chain is used for oxidative phosphorylation. Which compounds donate electrons to the electron- transport chain? H₂O NADH பப NAD+ ATP ADP FADH₂ FAD Which compounds are the final products of the electron-transport chain and oxidative phosphorylation? H₂O NADH NAD+ ΠΑΤΡ Π ADP FADH₂ FAD Which compound is the final electron acceptor in the electron-transport chain? Оно NADH NAD+ ATP ADP FADH₂ FADarrow_forwardHexokinase in red blood cells has a Michaelis constant (KM) of approximately 50 μM. Because life is hard enough as it is, let's assume that hexokinase displays Michaelis-Menten kinetics. What concentration of blood glucose yields an initial velocity (V) equal to 90% of the maximal velocity (Vmax)? [glucose] = What does the calculated substrate concentration at 90% Vmax tell you if normal blood glucose levels range between approximately 3.6 and 6.1 mM? Hexokinase operates near Vmax only when glucose levels are low. Hexokinase normally operates far below Vmax. Hexokinase operates near Vmax only when glucose levels are high. Hexokinase normally operates near Vmax mMarrow_forwardClassify each coenzyme or distinguishing characteristic based on whether it corresponds to catalytic or stoichiometric coenzymes. Catalytic coenzymes Answer Bank Stoichiometric coenzymes lipoic acid FAD used once coenzyme A regenerated thiamine pyrophosphate (TPP) NAD+arrow_forward
- The oxidation of malate by NAD+ to form oxaloacetate is a highly endergonic reaction under standard conditions. AG +29 kJ mol¹ (+7 kcal mol-¹) Malate + NAD+ oxaloacetate + NADH + H+ The reaction proceeds readily under physiological conditions. = Why does the reaction proceed readily as written under physiological conditions? The NADH produced during glycolysis drives the reaction in the direction of malate oxidation. The steady-state concentrations of the products are low compared with those of the substrates. The reaction is pushed forward by the energetically favorable oxidation of fumarate to malate. Endergonic reactions such as this occur spontaneously without the input of free energy. Assuming an [NAD+ ]/[NADH] ratio of 8, a temperature of 25°C, and a pH of 7, what is the lowest [malate]/[oxaloacetate] ratio at which oxaloacetate can be formed from malate? [malate] [oxaloacetate]arrow_forwardCalculate and compare the AG values for the oxidation of succinate by NAD+ and FAD. Use the data given in the table to find the E of the NAD+: NADH and fumarate:succinate couples, and assume that E for the enzyme-bound FAD: FADH2 redox couple is nearly +0.05 V. Oxidant Reductant " E' (V) NAD+ NADH + H+ 2 -0.32 Fumarate Succinate AG°' for the oxidation of succinate by NAD+: AG°' for the oxidation of succinate by FAD: 2 -0.03 Why is FAD rather than NAD+ the electron acceptor in the reaction catalyzed by succinate dehydrogenase? The electron-transport chain can regenerate FAD, but not NAD+. FAD is an oxidant, whereas NAD+ is a reductant. The oxidation of succinate requires two NAD+ molecules but only one FAD molecule. The oxidation of succinate by NAD+ is not thermodynamically feasible. kJ mol-1 kJ mol-1arrow_forwardUse the cellular respiration interactive to help you complete the passage. 2,4-dinitrophenol (DNP) was a popular ingredient in diet pills in the 1930s before it was discovered that moderate doses of the compound cause exceptionally high body temperature and even death. Complete the passage detailing how DNP's mechanism of action explains why it causes both high body temperature and weight loss. 2,4-dinitrophenol (DNP) causes of returning to the mitochondrial matrix through to pass directly across the inner mitochondrial membrane instead proteins. Because of DNP's effect on the mitochondrion, less energy is captured in the form of energy is instead wasted as heat. and more protons electrons ATP NADH sugars cytochrome ATP synthase heatarrow_forward
- To answer this question, you may reference the Metabolic Map. Select the reactions of glycolysis in which ATP is produced. 1,3-Bisphosphoglycerate 3-phosphoglycerate Glyceraldehyde 3-phosphate 1,3-bisphosphoglycerate Fructose 6-phosphate fructose 1,6-bisphosphate Phosphoenolpyruvate pyruvate Glucose glucose 6-phosphate Suppose 17 glucose molecules enter glycolysis. Calculate the total number of inorganic phosphate (P) molecules required as well as the total number of pyruvate molecules produced. P required: pyruvate produced: molecules moleculesarrow_forwardSuppose a marathon runner depletes carbohydrate stores after a four-hour run. The runner's nutritionist suggests replenishing carbohydrate stores by eating carbohydrates. However, the runner is also concerned about weight loss and wants to know if fats can be directly converted into carbohydrates. How should the nutritionist respond to the runner? Yes, the glyoxylate cycle can be used to convert acetyl CoA into succinate, which can then be converted into carbohydrates. No, the two decarboxylation reactions of the citric acid cycle preclude the net conversion of acetyl CoA into carbohydrates. No, the citric acid cycle converts acetyl CoA into oxaloacetate, but there is no pathway to form glucose from oxaloacetate. Yes, pyruvate carboxylase can convert acetyl CoA into pyruvate, which can be used to form glucose through gluconeogenesis.arrow_forwardThe crossover technique can reveal the precise site of action of a respiratory-chain inhibitor. Britton Chance devised elegant spectroscopic methods for determining the proportions of the oxidized and reduced form of each carrier. This determination is feasible because the forms have distinctive absorption spectra, as illustrated in the graph for cytochrome c. Upon the addition of a new inhibitor to respiring mitochondria, the carriers between NADH and ubiquinol (QH2) become more reduced, and those between cytochrome c and O₂ become more oxidized. Where does your inhibitor act? Complex I Complex II Complex III Complex IV Absorbance coefficient (M-1 cm x 10-5) 10 1.0 0.5 400 Reduced Oxidized 500 Wavelength (nm) 600arrow_forward
- Why are the electrons carried by FADH2 not as energy rich as those carried by NADH? FADH2 carries fewer high-energy electrons than NADH. OFADH2 is less negatively charged than NADH. OFADH2 has a lower phosphoryl-transfer potential than NADH. FADH₂ has a lower reduction potential than NADH. What is the consequence of this difference? Electrons flow from NADH to FADH2 before they are transferred to O₂. Electron flow FADH₂ to O, results in the production of more ATP than does electron flow from NADH. Electron flow from FADH₂ to O, pumps fewer protons than does electron flow from NADH. Electron flow from FADH, to O, consumes more free energy than does electron flow from NADH. A simple equation relates the standard free-energy change, AG", to the change in reduction potential, AE. AG=-FAE Then represents the number of transferred electrons, and F is the Faraday constant with a value of 96.48 kJ mol¹ V-¹. Use the standard reduction potentials provided to determine the standard free energy…arrow_forwardMatch each enzyme with its description. catalyzes the formation of isocitrate synthesizes succinyl CoA generates malate generates ATP converts pyruvate into acetyl CoA converts pyruvate into oxaloacetate condenses oxaloacetate and acetyl CoA catalyzes the formation of oxaloacetate synthesizes fumarate catalyzes the formation of a-ketoglutarate Answer Bank succinate dehydrogenase a-ketoglutarate dehydrogenase aconitase fumarase citrate synthase malate dehydrogenase pyruvate carboxylase pyruvate dehydrogenase complex isocitrate dehydrogenase succinyl CoA synthetasearrow_forwardcoo ☐ CH2 coo Malonate Determine how the concentration of each citric acid cycle intermediate will change immediately after the addition of malonate. The concentration of citrate will The concentration of isocitrate will The concentration of α-ketoglutarate will The concentration of succinyl CoA will The concentration of succinate will The concentration of fumarate will The concentration of malate will The concentration of oxaloacetate will Why is malonate not a substrate for succinate dehydrogenase? Malonate lacks a thioester bond that has high transfer potential. Malonate has two carboxylic acid groups. Malonate is not large enough to bind to the enzyme. Malonate only has one methylene group.arrow_forward
 BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
